Genome of Sezary syndrome points to potential treatment targets
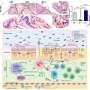
A genomic analysis of 37 patients with Sézary syndrome, a rare form of T-cell lymphoma that affects the skin and causes large numbers of atypical T-lymphocytes (an immune system disease) to circulate, reveals mutations in genes that affect T-cell signaling and those that interfere with cell cycle checkpoints that govern cell division, said researchers from Baylor College of Medicine and The University of Texas MD Anderson Cancer Center in a report in the journal Nature Genetics.
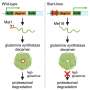
A key finding were activating mutations in genes called CCR4 and CARD11 in one-third of the patients. Already, patient studies are ongoing with experimental drugs that inhibit the CCR4 mutation, said Dr. David Wheeler, professor in the Baylor College of Medicine Human Genome Sequencing Center and a corresponding author of the paper. The studies are being carried out by Dr. Madeleine Duvic of MD Anderson, another corresponding author.
"These kinds of studies are taking us to the doorstep of personal genomics," said Wheeler, who leads the cancer genomics studies in the Human Genome Sequencing Center at Baylor. "We are finding particular treatment targets in some of these patients – targets for which we already have drugs or for which we can develop them."
"This research is important because it identifies genes that may be important in this rare cancer," said Dr. Linghua Wang, assistant professor of molecular and human genetics at Baylor and lead author of the study.
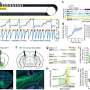
ZEB1, a gene essential for the differentiation of T-cells, was deleted in more than half the patients and two other immune system genes – IL32 and IL2RG – were overexpressed in nearly all cases. When the scientists analyzed two T-cell receptors (Vβ and Vα) and how they are expressed, they found that they were rearranged when the malignant T-cell clone expanded in one-third of patients.
More information: Linghua Wang et al. Genomic profiling of Sézary syndrome identifies alterations of key T cell signaling and differentiation genes, Nature Genetics (2015). DOI: 10.1038/ng.3444