SNP location helps predict disease aetiology
Nature Review Genetics has highlighted work by Milner Centre PhD student XianMing Wu.
Co-authored by Professor Laurence Hurst, the study concerns locations of disease-causing mutations in genes. The results have implications for diagnosis of genetic diseases.
Challenging a common belief
Genes have sections that are read into protein (exons) and sections that are thrown away (introns).
A common assumption is that disease-causing mutations in exons change the protein slightly. This would suggest that disease-associated mutations will be found equally at all positions in an exon. The team showed that this was not true.
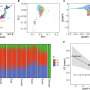
Using a database of thousands of mutations known to factor in human genetic diseases, they found that disease-causing mutations are much more likely to occur at the ends of exons.
XianMing commented: "The bias is really strong. The centres of our exons have only about half the number that we would have expected were the mutations randomly spread out."
Laurence added: "Over 80% of all disease causing mutations happen at these end parts. Our result suggests that a very high proportion of mutations are not causing disease by the mechanisms that people have typically assumed must be operating.
"We estimate that between a quarter and half of all disease-causing mutations act via this misprocessing route."
Previously overlooked significance
The team also discovered that some exons are better able to resist mutations. The team suggest the concept of the fragile exon, one in which one mutation can disrupt the exon finding machinery.
"One of the major consequences of our finding is that as a community we have probably been overlooking many really important mutations," said Laurence. "Just because we didn't see why they should be important."
More information: Liesbet Lieben. Disease genetics: SNP location helps predict disease aetiology, Nature Reviews Genetics (2015). DOI: 10.1038/nrg.2015.12