December 30, 2015 report
New embryo analysis technique helps screen out genetic problems prior to IVF
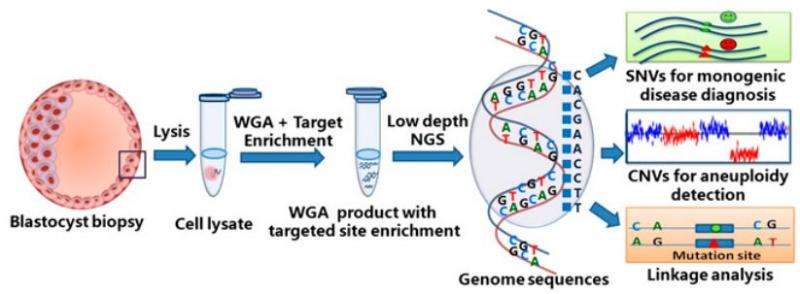
(MedicalXpress)—A new technique developed by a team of researchers affiliated with a number of facilities in China allows medical practitioners involved in IVF treatment to more easily weed out embryos with genetic defects prior to fertilization and implantation of a zygote into a host uterus. In their paper published in Proceedings of the National Academy of Sciences, the team describes the new technique and the circumstances under which it is useful.
Couples having trouble conceiving a child turn to medical specialists to help them overcome whatever obstacles they may be facing—in some cases one option is to provide in-vitro fertilization, where an egg is surgically removed from the mother-to-be, and sperm is obtained from the father to be (or a stand-in), and the results are mixed in a lab, and if the egg becomes fertilized, it is implanted into the womb of the mother, or a suitable stand-in. Unfortunately, in some cases the process doesn't always go according to plan and the couple winds up with no baby—the odds diminish as the age of the mother donating the egg reaches over forty, due do what is known as a chromosomal aneuploidy, where the number of chromosomes is not normal. Other problems come about due to gene mutations from either parent or when a gene is missing.
Over the years, scientists have developed different tests to check for aneuploidy or gene mutations, but until now, there has been no test to check for both at one time, which is important, because testing can be hard on an embryo, causing other problems. In this new effort, the researchers have come up with a way to test for both, a technique called mutated allele revealed by sequencing with aneuploidy and linkage analyses—it is a next generation sequencing method for single-cell genome amplification that involves multiple heating/cooling and looped-based amplification cycles.
The technique has been tried successfully on two different couples already, one where the father had a gene abnormality that caused bone spurs, and the other, where the mother had a gene mutation that caused skin and sweat problems. The researchers were able to successfully weed out the embryos that carried the gene mutations allowing both couples to have babies that were free of their parents' maladies.
More information: Liying Yan et al. Live births after simultaneous avoidance of monogenic diseases and chromosome abnormality by next-generation sequencing with linkage analyses, Proceedings of the National Academy of Sciences (2015). DOI: 10.1073/pnas.1523297113
Abstract
In vitro fertilization (IVF), preimplantation genetic diagnosis (PGD), and preimplantation genetic screening (PGS) help patients to select embryos free of monogenic diseases and aneuploidy (chromosome abnormality). Next-generation sequencing (NGS) methods, while experiencing a rapid cost reduction, have improved the precision of PGD/PGS. However, the precision of PGD has been limited by the false-positive and false-negative single-nucleotide variations (SNVs), which are not acceptable in IVF and can be circumvented by linkage analyses, such as short tandem repeats or karyomapping. It is noteworthy that existing methods of detecting SNV/copy number variation (CNV) and linkage analysis often require separate procedures for the same embryo. Here we report an NGS-based PGD/PGS procedure that can simultaneously detect a single-gene disorder and aneuploidy and is capable of linkage analysis in a cost-effective way. This method, called "mutated allele revealed by sequencing with aneuploidy and linkage analyses" (MARSALA), involves multiple annealing and looping-based amplification cycles (MALBAC) for single-cell whole-genome amplification. Aneuploidy is determined by CNVs, whereas SNVs associated with the monogenic diseases are detected by PCR amplification of the MALBAC product. The false-positive and -negative SNVs are avoided by an NGS-based linkage analysis. Two healthy babies, free of the monogenic diseases of their parents, were born after such embryo selection. The monogenic diseases originated from a single base mutation on the autosome and the X-chromosome of the disease-carrying father and mother, respectively.
© 2015 MedicalXpress