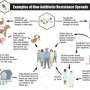
Since it first emerged more than half a century ago, a particular strain of multidrug-resistant Salmonella has spread all over the world. Now researchers have figured out why this strain, Salmonella Typhimuriam DT104, has been so successful. This new knowledge could prove valuable in combating other successful pathogens, according to the authors. The study is published ahead of print March 4th in Applied and Environmental Microbiology, a journal of the American Society for Microbiology.
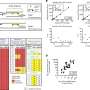
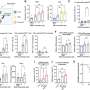
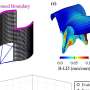
In order to construct the history of this strain, the investigators performed whole-genome sequencing of samples of DT104 that had been collected from patients over more than 40 years, from 1969 to 2012, in 21 countries, on six continents. Very tiny changes in the genome that took place over time enabled them to construct the strain's family tree (which scientists call a phylogenetic tree). The sequences have also made it easy to estimate roughly when the pathogen acquired the resistance genes.
DT104's success was due in no small part to its resistance to at least five antibiotics, including ampicillin, chloramphenicol, streptomycin, sulphonamide, and tetracycline, said corresponding author, Pimlapas Leekitcharoenphon, PhD.
Further abetting its spread, unlike other strains of DT Salmonella, DT104 was able to infect numerous livestock species, including cattle, poultry, pigs, and sheep, said Leekitcharoenphon. "Having multiple hosts increases the chances of dissemination," she explained. Leekitcharoenphon is a postdoctoral researcher at the Research Group for Genomic Epidemiology, National Food Institute, Technical University of Denmark, Lyngby.
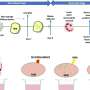
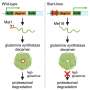
Using a program that took into account the rate of mutations in DT104, the investigators estimated that it first emerged in 1948 as an antibiotic susceptible pathogen. It is not clear exactly when DT104 first acquired the multidrug resistance-containing transposon. Transposons are mobile genetic elements that can carry antibiotic resistance genes, and that can jump from one genome to another. In the case of DT104, transposons have been identified as the sources of the resistance genes. The study suggests that the first acquisition of antibiotic resistance may have happened in 1972. However, multidrug-resistant DT104 was first reported in 1984 in the United Kingdom.
The new results also illuminated, for the first time, the results of a program in Denmark to eradicate all pigs infected with DT104, which had begun in 1996, but was stopped in 2000 due to financial pressures. It turns out that program was quite successful.
"If we know and understand the past, we might be able to solve the current resistance problems and prevent future ones," said Leekitcharoenphon.
Journal information: Applied and Environmental Microbiology
Provided by American Society for Microbiology