Three lessons gut microbes have taught us about antibiotics
Antibiotics have proven to be a double-edged sword: capable of killing a range of bacteria that cause infections, but also depleting our gut microbes, impairing our immune system, and increasing vulnerability to infection by superbugs. The lessons learned from how antibiotics impact the body, both positively and negatively, are identifying new approaches to prevent and/or correct the adverse side effects on our "good" gut bacteria, say authors of a Review published May 10 in Trends in Molecular Medicine.
Lesson #1. Antibiotics disrupt the communication between gut microbes and the immune system, creating conditions that favor infection.
In the last 10 years, researchers have established that "good" commensal bacteria lining our intestines help stabilize the immune system through molecular crosstalk with immune cells. Add antibiotics, and many of the signals our microbes send are lost, temporarily disrupting the ability of immune cells to function normally.
Disturbances to the human microbiota early in life have been linked to immune or metabolic difficulties, such as asthma and increased body mass, which can persist into adulthood. Evidence suggests that there are key moments when the presence of certain commensal bacteria is necessary for immune development. "Treatment with antibiotics, by reducing the amount of competing commensals, increases available sugars and, in some settings, favors pathogen expansion," says senior author Eric Pamer, a physician-scientist at the Memorial Sloan Kettering Cancer Center.
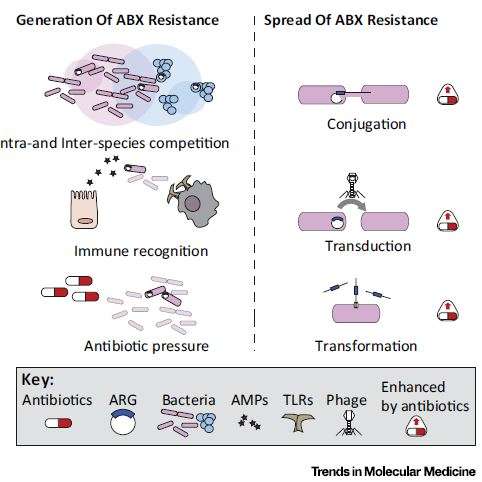
Lesson #2. Antibiotic use becomes a selective pressure on bacteria, driving antibiotic resistance.
Antibiotic resistance genes are naturally present in bacterial communities found anywhere from permafrost to the bacteria in our intestines. One problem with antibiotics is that by killing populations without such genes, those that survive multiply and transfer antibiotic resistance genes to other bacterial groups.
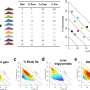
Recent research has shown that the pool of antibiotic resistance genes within an individual's microbiome "expands over time" as the person ages and is exposed to more antibiotics. One surprising study (DOI: 10.1093/gbe/evv102) demonstrated that 40% of bacteria from human and animal hosts carried resistance genes to one of the categories of broad-spectrum antibiotics (quinolones), "even in subjects that had never been exposed clinically," notes first author Simone Becattini, a research laboratory fellow in Pamer's lab.
Lesson #3. Antibiotics aren't the only solution, and strategies to specifically target infectious bacteria are in early development.
Future approaches to treating bacterial infections will either complement current antibiotics or provide a more narrowly targeted alternative. One complementary treatment boosts the production of antimicrobial factors by immune cells; following bacterial eradication with antibiotic treatment, the injection of immune-stimulatory molecules into the blood jolts immune cells and has been shown to successfully protect mice from potential bacterial colonization in the intestine.
Promising alternatives to broad-spectrum antibiotics include the following:
- Bacteriocins, which are proteins produced by bacteria to kill competitors. "The use of bacteriocins or bacterial strains that are engineered to carry bacteriocins would selectively kill the pathogens without altering the microbiota," says Becattini.
- CRISPR-CAS9 gene editing can be used to cut out antibiotic resistant genes from infectious bacteria. This approach has already been shown to efficiently target specific bacteria where signs of resistance fail to develop afterwards.
- Fecal material transplants can also help restore commensal gut communities, mucus production, antimicrobial peptide secretion, and provide colonization resistance against pathogens that are cleared or can no longer expand.
These approaches are currently in various states of clinical development, and researchers are closely watching how they perform.
More information: Trends in Molecular Medicine, Becattini et al.: "Antibiotic-induced changes in the intestinal microbiota and disease" www.cell.com/trends/molecular- … 1471-4914(16)30007-7 , DOI: 10.1016/j.molmed.2016.04.003