Gene signature in ovarian cancer predicts survival and offers new drug target

A new UK study has identified a gene signature that predicts poor survival from ovarian cancer. The study also identified genes which help the cancer develop resistance to chemotherapy - offering a new route to help tackle the disease.
The study, published in the International Journal of Cancer, examined the role of HOX genes in ovarian cancer resistance and whether a drug known as HXR9 which targets HOX, could help prevent the resistance from developing.
The HOX gene family enables the remarkably rapid cell division seen in growing embryos. Most of these genes are switched off in adults, but previous research has shown that in several cancers, including ovarian cancer, HOX genes are switched back on, helping the cancer cells to proliferate and survive.
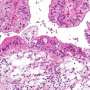
Led by Professor Richard Morgan, Director of the University of Bradford's Institute of Cancer Therapeutics, researchers analysed tissue samples from 99 women with epithelial ovarian cancer (EOC) - the most common form - and compared these with healthy ovarian and fallopian tube tissue samples.
Little to no HOX expression was found in normal ovarian tissue whereas 36 of the 39 HOX genes were found at high levels in tissue samples of the EOC subtype known as 'high grade serous', which accounts for approximately 80% of epithelial ovarian cancers. A strong five-gene signature was found in all patients who succumbed to the disease, irrespective of their length of survival.
The team also conducted extensive tests on cells and preliminary tests on mice using HXR9 - a peptide drug developed by Professor Morgan which blocks the function of the proteins expressed by HOX genes, forcing cancer cells to close down and die.
The team tested both HXR9 and cisplatin, the most common drug currently used to treat ovarian cancer, and a combination of the two. They found that combining the two drugs significantly increased the number of cancer cells which were killed compared to either drug used alone.
Co-author Dr Zoe Kelly, who carried out this research at the University of Surrey, said: "We've identified a set of genes which play a contributing role in resistance to chemotherapy, which is a major problem in the treatment of ovarian cancer. We also have strong and extensive cell line data which shows that using HXR9 can overcome this drug resistance, making the cell more susceptible to chemotherapy treatment.
She added: "The results in mice were encouraging, but more muted: treated mice survived for longer, but the cell killing of the combination approach was only marginally better than HXR9 used on its own. However, these tests were carried out over a very short timeframe, and I believe that more extensive tests in the mouse model would show clearer results. This needs to be the next step for this research."
Professor Morgan said: "This is the first comprehensive analysis of HOX gene expression in ovarian cancer and the first study to analyse changes in HOX expression in resistant cancer cells. The results strongly suggest that targeting these genes as a new treatment approach warrants further investigation. It also supports our belief that HXR9 should be further developed and tested in clinical trials."
More information: Zoe Kelly et al, The prognostic significance of specificgene expression patterns in ovarian cancer, International Journal of Cancer (2016). DOI: 10.1002/ijc.30204