April 20, 2017 report
Using RNA sequencing to diagnose patients with rare muscle conditions

(Medical Xpress)—An international team of researchers has developed a way to use RNA sequencing to help in diagnosing patients with rare genetic muscle conditions. In their paper published in the journal Science Translational Medicine, the group describes their technique, how it works, and how effective they found it when testing actual patients.
Muscular dystrophy is a group of diseases of the muscle that are thought to have a genetic cause. Muscles weaken to different degrees and in different part of the body depending on type, and in some cases, cease to work at all. There are nine major categories that include 32 types, many of which are difficult to diagnose. None are curable, though there are many treatments.
As the researchers note, genome and exome sequencing has proved to be an effective means for diagnosing rare genetic disorders, including those of the muscle, but it is still not good enough. Success rates for MD vary from 25 to 50 percent. To improve those percentages, the researchers with this new effort took another approach—using RNA transcriptome sequencing as a complementary tool.
The technique is based on the fact that different tissues in the body do not express all of the genes that exist within a given genome, which means that at times, DNA sequences do not alter the RNA and the proteins that cells synthesize when performing various functions.
The group performed their RNA synthesis technique on muscle biopsies taken from 50 patients who each had a genetic ailment that could not be identified through genome or exome sequencing. They report that they were able to identify 17 cases of muscular dystrophies, which worked out to be a rate of 35 percent—a major improvement. They note that the diagnosis in several of those cases was extremely difficult to detect using other methods.
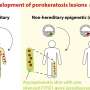
The researchers also report that they found one new mutation that could be linked to a certain kind of muscular dystrophy, and found it in 27 people from a different group of undiagnosed patients. That variant, they also note, does not appear in the 1000 Genomes Project database, and the researchers suggest it might be present in approximately 25 percent of undiagnosed dystrophies.
More information: Beryl B. Cummings et al. Improving genetic diagnosis in Mendelian disease with transcriptome sequencing, Science Translational Medicine (2017). DOI: 10.1126/scitranslmed.aal5209
Abstract
Exome and whole-genome sequencing are becoming increasingly routine approaches in Mendelian disease diagnosis. Despite their success, the current diagnostic rate for genomic analyses across a variety of rare diseases is approximately 25 to 50%. We explore the utility of transcriptome sequencing [RNA sequencing (RNA-seq)] as a complementary diagnostic tool in a cohort of 50 patients with genetically undiagnosed rare muscle disorders. We describe an integrated approach to analyze patient muscle RNA-seq, leveraging an analysis framework focused on the detection of transcript-level changes that are unique to the patient compared to more than 180 control skeletal muscle samples. We demonstrate the power of RNA-seq to validate candidate splice-disrupting mutations and to identify splice-altering variants in both exonic and deep intronic regions, yielding an overall diagnosis rate of 35%. We also report the discovery of a highly recurrent de novo intronic mutation in COL6A1 that results in a dominantly acting splice-gain event, disrupting the critical glycine repeat motif of the triple helical domain. We identify this pathogenic variant in a total of 27 genetically unsolved patients in an external collagen VI–like dystrophy cohort, thus explaining approximately 25% of patients clinically suggestive of having collagen VI dystrophy in whom prior genetic analysis is negative. Overall, this study represents a large systematic application of transcriptome sequencing to rare disease diagnosis and highlights its utility for the detection and interpretation of variants missed by current standard diagnostic approaches.
© 2017 Medical Xpress