Environmental pressures on opportunistic fungal pathogen
With an estimated one million cases diagnosed worldwide each year, the human pathogen Cryptococcus neoformans, which can cause life-threatening fungal infections in immunocompromised patients, is an important health concern. In a study published today in Genome Research, scientists identified natural genomic variation in C. neoformans that may influence prevalence and disease severity.
Researchers from the Broad Institute of MIT and Harvard, Duke University Medical Center, and elsewhere sequenced 387 environmental and clinical isolates of C. neoformans predominantly from sub-Saharan Africa, where the disease burden of cryptococcosis is high. Because the pathogen is environmentally acquired and cannot be transmitted human-to-human, any genetic variation leading to changes in virulence must be coincident with other selective pressures.
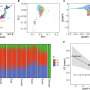
Using whole-genome sequencing data, the researchers found approximately 1 million variants across samples, and were able to clearly differentiate the three known C. neoformans lineages, VNI, VNII, and VNB. Additionally, VNB could be further subdivided into two previously suggested subgroups, VNBI and VNBII, based on multiple approaches. Notably, the two subgroups showed vastly different frequencies in the environment, with VNBII almost exclusively found in clinical samples. Large scale phenotypic profiling revealed that VNBII clinical isolates are less pigmented, which protects the fungus from environmental stress, and also less resistant to oxidative damage. After characterizing recombination levels in these subgroups, the researchers carried out genome-wide association studies and scans for selective signatures, which identified specific variants associated with clinical isolates including some linked to known virulence factors.
"Our work begins to examine how natural variation contributes to differences in human infection and suggests that some genetic pathways are less important during human infection than for growth in the environment," said corresponding author and Senior Group Leader, Fungal Genomics at the Broad Institute, Christina Cuomo said.
More information: Genome Research (2017). DOI: 10.1101/gr.218727.116