Next-generation metagenomics sequencing may sleuth out hard-to-find viruses in the blood
Next-generation metagenomics sequencing (NGMS) may have broad potential for discovering new viral infections in the blood, such as human hepegivirus-1 (HHpgV-1), that currently cannot be found by conventional methods. Whether or not HHpgV-1 causes human disease remains to be seen. The findings are published in Annals of Internal Medicine.
Humans are teeming with microbes that contribute to health and disease. NGMS has been used in clinical practice to detect unsuspected pathogens and also to characterize the composition of complex populations of recognized viruses, such as HIV-1 and hepatitis C virus (HCV). While NGMS can uncover novel sequences, the limits of detection are not clearly defined. Understanding the sensitivity and accuracy of NGMS compared with quantitative clinical standards is critical to determining its clinical application.
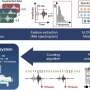
Researchers at Johns Hopkins School of Medicine studied plasma samples from persons who inject drugs co-infected with HIV and HCV who were enrolled in a prospective study of HCV dynamics after pegylated interferon-?2b (IFN) administration and before and after antiretroviral therapy (ART). This is important because persons who inject drugs have higher risk of blood born infections and thus can be "sentinels" of emerging blood borne diseases. The researchers compared viral nucleic acid in plasma by NGMS and quantitative polymerase chain reaction (PCR).
The researchers found that NGMS was insensitive for detection of viruses with relatively low plasma nucleic acid concentrations, but was able to discover an unexpected, novel hepegivirus in the human virome, HHpgV-1. Furthermore, the researchers found HHpgV-1 sequences in liver tissue. The significance of this finding is not yet known, but it sheds light on the sensitivity and accuracy of NGMS for detecting viruses in the blood.
More information: Annals of Internal Medicine (2017). http://annals.org/aim/article/doi/10.7326/M17-0085