Researchers grow micro-tumour models for more targeted cancer therapy
Researchers have successfully grown micro-tumour models outside the human body and used them to identify alternative treatment options for cancer patients who are suffering from resistant and/or metastatic disease, meaning that they fail to respond to standard-of-care cancer drugs.
This approach shows the potential to predict treatment response to drugs and help clinicians identify appropriate therapeutic options for cancer patients. The study, published in Nature Communications, was jointly led by A*STAR's Genome Institute of Singapore (GIS) and National Cancer Centre Singapore (NCCS).
The study involved growing the patients' own tumour material ex vivo in a dish as "micro-tumour avatars", with the team successfully generating 24 patient-derived "avatar-models". These micro-tumour models were then screened for therapeutic and genetic vulnerabilities against a panel of clinically relevant drugs, including standard-of-care treatments such as chemotherapy, radiotherapy or targeted anti-cancer drugs. This approach is known as phenotype-based treatment.
In addition, genomic characterisation of these models helped identify new biomarkers that could predict tumour progression as well as the patients' response to different classes of drugs.
Among the group of patients involved in the study, two of them went on to receive the phenotype-based treatment in the clinic, and responded remarkably well to the treatment options that were identified, including one where the tumour volume shrank by more than 95% within six weeks of treatment, and the other where the expression of a cancer antigen marker in the blood was reduced by over 90%.
The researchers plan to subsequently scale this effort to reveal specific predictive gene expression and mutation signatures that could be used to stratify different patient groups and corresponding treatment options. This would help to guide future treatment strategies validated through larger clinical trials.
"Our approach is unique in the sense that we are using phenotype-driven approaches prospectively to define real-time clinical management of disease, and then investigating the genomic basis for response to identify novel prognostic biomarkers. The ultimate goal would be for this innovative strategy to be available to every cancer patient in future," explained the study's lead author Dr Ramanuj DasGupta, Group Leader, Cancer Therapeutics and Stratified Oncology at the GIS.
Dr DasGupta added, "To truly assess what treatments may or may not work for a given patient, we need to let the patient's own cancer cells tell us the genetic basis for their response or lack thereof to certain treatments, instead of using historically existing cell lines. This approach would be exceptionally useful where there is a lack of treatment options, for instance in late-stage patients, or a lack of clear biomarker-driven treatment strategies."
"To get this right takes a lot more than just the work reflected in this paper alone. There were years of struggles on both sides to get protocols in place to be able to recruit patients, grow the tumours outside of the patient's body, testing for genetic and therapeutic vulnerabilities at the screening setup at GIS1, and have it run like clockwork, with data analysis in a meaningful manner, making sense of the results, bringing that information back to the hospital for discussions, and finally treating the patient again. Each of these steps is herculean to say the least but this is a small step towards the future where treatment for any patient will be based on their unique characteristics and not assumptions from group studies based on other populations. This takes planning, determination, mutual respect for each other's strength and a dedication to see this through," said Dr Gopal Iyer, co-lead author of the study and Senior Consultant Surgical Oncologist at the NCCS.
Executive Director of GIS, Prof Ng Huck Hui said, "Tumour heterogeneity poses a major challenge in cancer therapies because different clonal populations of cells within a tumour can respond very differently to standard-of-care treatments, often resulting in treatment failure or relapse for unknown reasons. I am delighted this multi-institutional collaborative effort has resulted in a novel approach that can help us advance precision medicine."
Cancer remains a major cause for disease-related mortality worldwide. It accounted for 8.8 million deaths in 2015; nearly 1 in 6 global deaths was due to cancer2. 70% of all cancer deaths afflict people in the developing world, including countries in Africa, Asia, Central and South America.
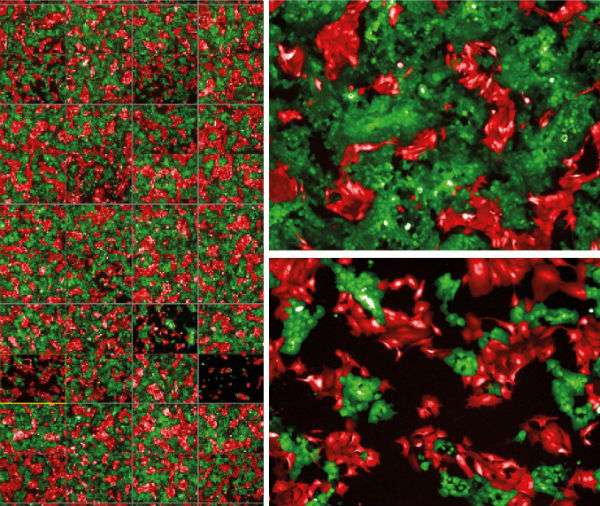
Personalised screening for drug sensitivity in patient-derived models. Green cells represent the primary tumour cells while the red cells represent metastatic tumours. The different micrographs represent the effect of drug treatment on primary versus metastatic tumour models derived from the same patient.
The research findings described in this media release can be found in the scientific journal Nature Communications, under the title, "Phenotype-driven precision oncology as a guide for clinical decisions one patient at a time."
More information: Shumei Chia et al. Phenotype-driven precision oncology as a guide for clinical decisions one patient at a time, Nature Communications (2017). DOI: 10.1038/s41467-017-00451-5