Improving prediction accuracy of Crohn's disease based on repeated fecal sampling

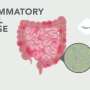
Researchers at the University of California San Diego Center for Microbiome Innovation (CMI) have found that sampling the gut microbiome over time can provide insights that are not available with a single time point. The findings could help doctors and researchers more accurately determine if a patient has Crohn's disease. The findings were published as a letter in Gut on October 21, 2017.
Researchers say the idea for the work came from a recently published study on the instability of the gut microbiomes of patients with Crohn's disease. "It is difficult to get a useful picture by collecting one fecal sample, and this property is likely what hinders our ability to do so," said lead author on the study and researcher in Center director Rob Knight's lab Yoshiki Vázquez-Baeza.
According to Vázquez-Baeza, we all have ever-changing microbiomes, but people with Crohn's Disease appear to have microbiomes that change much more frequently. He and his team wondered, could sampling the microbiome over time provide a new way to classify the disease? Furthermore, could a machine-learning model use this increased variability as a 'tell-tale' to discriminate between affected and unaffected subjects?
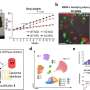
In order to investigate these questions, the researchers collected stool samples daily from a total of 31 people. According to Knight , also a professor in the Departments of Pediatrics and Computer Science and Engineering at UC San Diego, the researchers believe this is the most densely sampled longitudinal study of Crohn's disease.
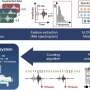
Following sample collection, the researchers created a computational model to gain insights from the data. When multiple fecal samples per subject were used, the model was able to predict whether someone had Crohn's disease better even than biopsy samples, which are more expensive and inconvenient to collect.
The methodology was repeated and the results replicated in a second cohort. According to Vazquez-Baeza, the results highlight the importance of treating Crohn's Disease as a volatile, time-varying condition, even during clinical remission.
What's next? "Testing this in a larger cohort would be a wonderful next step," said Vázquez-Baeza. "We're hoping to see whether this is robust to fairly heterogeneous populations, and if the features themselves are consistent."
More information: Yoshiki Vázquez-Baeza et al. Guiding longitudinal sampling in IBD cohorts, Gut (2017). DOI: 10.1136/gutjnl-2017-315352