New resistance mechanism in the often multidrug resistant pathogen Acinetobacter baumannii
A team of Australian and Portuguese investigators has discovered yet another resistance mechanism in the pathogen, Acinetobacter baumannii, in this case, one that blocks the critical antibiotic-of-last-resort, colistin. A. baumannii is a highly troublesome pathogen globally, infecting primarily patients in intensive care units with ventilator-associated pneumonia, blood stream infections, and urinary tract infections. The research is published in Antimicrobial Agents and Chemotherapy, a journal of the American Society for Microbiology.
The genesis of the research was the discovery by Portuguese clinicians that A. baumannii from a bloodstream infection remained in an elderly patient even after treatment. "Colistin was being used as a last-resort treatment as the strain was highly multidrug resistant," the patient having already been treated with six different antibiotics, said corresponding author John Boyce, PhD, BSc (Hons).
"The infection could not be cleared as the strain developed high-level colistin resistance during treatment," said Dr. Boyce, who is Associate Professor, Infection and Immunity Program, Monash Biomedicine Discovery Institute, and Department of Microbiology, Monash University, Melbourne, Australia. "The clinicians wanted to understand how the strain had become highly colistin resistant, so we determined whole genome sequences of the pre- and post-treatment isolates." (Dr. Boyce's conversations with these clinicians at a conference in Rome led to the collaboration on this research.)
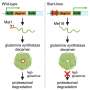
The investigators then compared those sequences to identify the genetic changes associated with the acquisition of high-level colistin resistance. They found only one difference: a gene called hns had been disrupted by an insertion sequence element (a piece of DNA that can insert itself into a chromosome), preventing normal expression. The hns gene encodes a protein that regulates gene transcription. Disrupting such a transcription regulator would change expression of many other genes in the bacteria.
To determine how disruption of the regulatory gene, hns affected colistin resistance, the researchers compared transcription products of genes from pre- and post-treatment isolates. Expression of a gene that can boost colistin resistance had increased post-treatment.
Dr. Boyce and his collaborators performed several additional experiments that confirmed that the disruption of hns was responsible for the resistance, including inactivating hns in the non-resistant strain, which caused that strain to become "highly colistin resistant," said Dr. Boyce.
"Colistin is a 'last resort' antibiotic that is generally used to treat infections caused by bacteria that are resistant to all other antibiotics," said Dr. Boyce. "Development of colistin resistance in strains that are already multidrug resistant is incredibly alarming; the threat of pan-resistant infections is very real... development of high-level colistin resistance in this instance played a major role in treatment failure and led to an infection that could not be resolved (although the patient had other comorbidities)."
"Understanding the mechanisms behind development of colistin resistance in clinical settings can allow us to identify treatments that minimize or circumvent resistance development, or will in future allow us to develop drugs that target the regulatory systems that control resistance," said Dr. Boyce.