Quick identification of multidrug-resistant pathogens

If doctors diagnose a patient with blood poisoning, the patient will immediately be administered a broad-spectrum antibiotic. In many cases, however, the drug is ineffective. Multidrug-resistant pathogens are often the reason why sepsis spreads through the body, ultimately resulting in the patient's death. As things stand, antibiotic resistance tests can take several days. In the PathoSept project, Fraunhofer researchers and partners are developing an end-to-end modular system that reduces the time needed to identify antibiotic-resistant pathogens to nine hours.
According to the World Health Organization (WHO), infections with multidrug-resistant pathogens pose one of the greatest risks to human health. And such pathogens are on the rise – most significantly in hospitals. Blood poisoning is one of the most severe types of infection. In Germany alone, more than 56,000 people die from sepsis each year. Given that the criticality of the infection increases by the hour, anyone who contracts it must be treated as quickly as possible. But, in many cases, the broad-spectrum antibiotic given to patients is ineffective because the bacteria have developed a resistance to the drug. As things stand, it can take up to five days for the right treatment to be administered following the first tentative diagnosis. The problem lies in the fact that time-consuming culture tests are required to identify resistant bacteria: after a blood sample has been taken, the bacteria must be left to multiply before the analysis can begin.
Initiating targeted treatment after nine hours
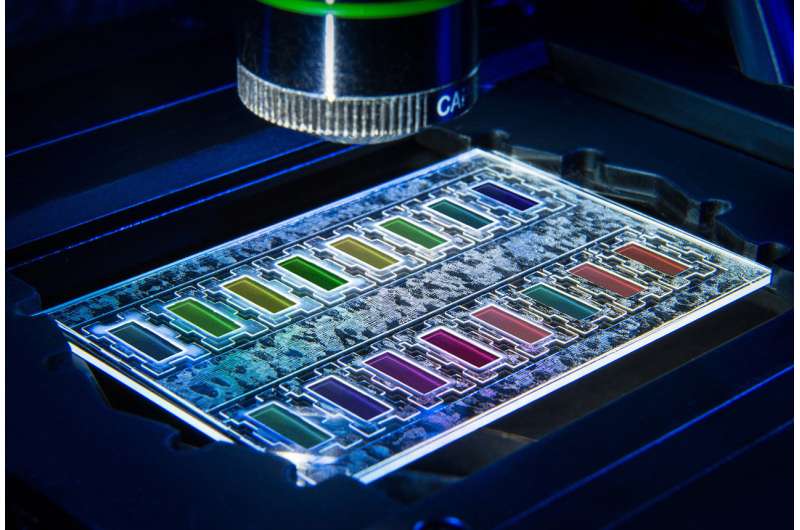
A new, modular system is set to speed up this process significantly. The first successful tests show that, in the future, doctors will be able initiate targeted treatment after just nine hours, since both the bacterium causing the infection and the right antibiotic to fight it can be identified in this time. In the PathoSept project (see box), researchers from the Fraunhofer Institute for Applied Information Technology FIT and several other partners developed a chip that can be used to analyze the growth behavior of bacteria under the influence of antibiotics. What sets this project apart is that the partners combined different methods to minimize the duration of the analysis while also optimizing cost efficiency. Fouad Bitti, a researcher at Fraunhofer FIT, explains the rationale: "Due to phenotypic and genotypic variability, no single diagnostic method today is capable of delivering results that are 100 percent reliable."
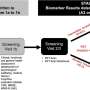
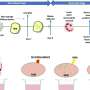
This system for detecting multidrug-resistant pathogens comprises four modules: In addition to a cultivation module for the controlled multiplication of pathogens, it contains protocols for separating the pathogens as well as highly sensitive assays for identifying the pathogens using qPCR – a nucleic acid amplification method based on the conventional polymerase chain reaction (PCR) principle. The core component of the system, a growth monitor that quickly quantifies resistances, was developed by researchers at Fraunhofer FIT, as was the cultivation module.
Software-supported resistance diagnostics
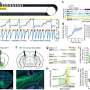
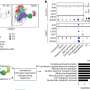
In the PathoSept project, pathogens are grown in the cultivation module to a critical limit and then added to 96 jars containing an antibiotic and nutrient solution. Featuring all the requisite analysis software, the growth monitor tracks and documents how the pathogens develop in real time. Algorithms evaluate the images recorded of the bacteria and extrapolate the growth curve. In this way, it is possible to determine after just a few hours whether the given drug is effective or whether the bacteria are resistant to it and will thus continue to multiply. Using standard software for flexible integration, the growth monitor calculates how the pathogens will develop over a longer time frame. The program analyzes both the size of the bacterial population – from which the exact number of bacteria can be deduced – and the ratio of living bacteria to those that have been killed. This enables the researchers to see which antibiotic kills the pathogens the quickest, the concentrations required and which bacteria have become resistant to the drug. "This allows us to initiate targeted treatment, which is a major advantage when you consider that doctors today have to administer a cocktail of antibiotics in the hope that one of them will be effective," says Bitti.
The diagnostic system is a bench-top device that is suitable for use in all medical laboratories. Thanks to its standard protocols and interfaces, it can easily be integrated into existing systems. The growth monitor is currently undergoing clinical testing in Germany, at the university clinics in Aachen and Bonn. Evaluation of the trials will then pave the way for the first demonstrator product. Clinical tests are also underway on the cultivation module. Bitti concludes: "In the future, our end-to-end modular diagnostic system will drastically reduce the mortality rate of sepsis patients."