Researchers report connection between intestinal bacteria and development of diabetes
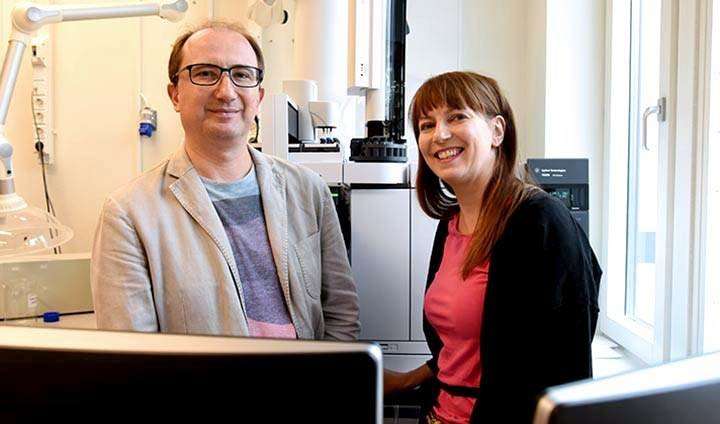

Researchers at Örebro University have, together with a well-known research team in Denmark, developed a method for studying how metabolism in gut bacteria influences health. Their method will now be published in its entirety in the scientific journal Nature Protocols.
The method, which researchers Tuulia Hyötyläinen and Matej Orešič have worked on and developed for more than ten years, is used for metabolomics studies—where information about thousands of molecules involved in the cells' metabolism is retrieved by means of chemical analysis. Using this method, researchers have been able to, among other things, study the connection between metabolism in gut bacteria and the development of diabetes, a study which was published in Nature 2016.
"But the method may be used by all researchers wanting to apply metabolomics in their studies," says Matej Orešič, researcher in medicine at Örebro University.
Scientific methods as important as results
By applying this method used by the Örebro researchers, it is possible to analyse some 2,000 metabolites from one blood sample. Metabolites are microscopic molecules, for example amino acids, lipids and sugar molecules, formed as a result of metabolism.
"Collecting data is an important part of the analysis but doesn't always require as much time as the actual data analysis. That's when this large amount of collected data will need to be linked to with biological and medical questions," says Tuulia Hyötyläinen, Professor of Chemistry at Örebro University.
In the published article, the working method protocol is described in detail. As is often the case, the method section of a scientific article is difficult for other researchers to reproduce, especially in complex studies with extensive data quantities.
"Scientific methods are as important as research results. It is important to use reliable methods to produce data of high quality," explains Matej Orešič.
Preventing gluten intolerance
The research duo is now working on identifying how metabolism in young children can affect the development of gluten intolerance later in life.
"We have seen changes in the metabolism of lipids in babies before they have been exposed to gluten via their diet. This discovery can perhaps lead to a better understanding of the development of gluten intolerance and perhaps help us to prevent the disease," adds Matej Orešič.
They are also investigating the relationship between exposure to various environmental pollutants in early life and the development of type 1 diabetes.
"We are trying to find biomarkers which may be used in the diagnosis of various diseases and at an earlier stage than is currently possible. It is also interesting to study which metabolites that are produced by gut bacteria and how these then influence our metabolism," says Tuulia Hyötyläinen.
"We are working continuously on improving the entire analysis chain, from taking specimens to data analysis. And now other researchers may also use our method," adds Matej Orešič.
More information: Helle Krogh Pedersen et al. A computational framework to integrate high-throughput '-omics' datasets for the identification of potential mechanistic links, Nature Protocols (2018). DOI: 10.1038/s41596-018-0064-z
Tuulia Hyötyläinen et al. Genome-scale study reveals reduced metabolic adaptability in patients with non-alcoholic fatty liver disease, Nature Communications (2016). DOI: 10.1038/ncomms9994