Interregional differences in somatic genetic landscape diversify prognosis in glioblastoma

Researchers from Osaka University, in cooperation with the Institute for Clinical Research and medical institutes participating in the Kansai Molecular Diagnosis Network for CNS Tumors (KNBTG), conducted the largest-ever retrospective cohort study for Japanese patients with glioblastoma (GBM), proposing an underlying prognosis biomarker responsible for the survival difference between two cohorts: an original Japanese cohort and a dataset from The Cancer Genome Atlas (TCGA). Their research results were published in Acta Neuropathologica Communications.
Clinical outcome of GBM has been reported to vary by region, and patients from Asia in particular have a better prognosis than those from any other region independent of treatment and social background. However, the cause of interregional survival differences in GBM remains poorly understood.
To unveil regional survival differences, the research team conducted genetic analysis of Japanese patients with the primary IDH-wildtype GBM registered in KNBTG. They investigated 140 KNBTG cases and 152 TCGA cases, both of which had been treated by current standard therapy (surgery followed by chemoradiation therapy), to conduct exploration of prognostic biomarkers and survival analysis.
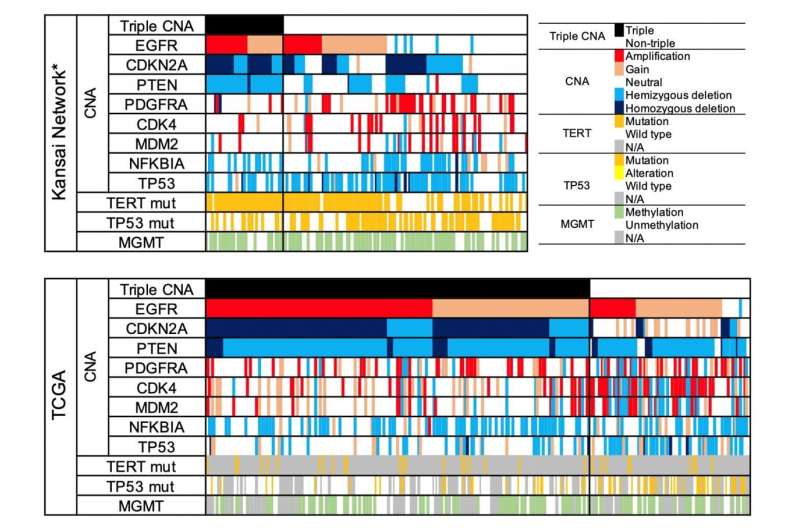
As a result, there were molecular disproportions between these two cohorts; that is, there was unequal distribution of specific copy number alterations (CNAs), such as EGFR gain, CDKN2A deletion, and PTEN deletion. They found that the triple overlap CNAs in these loci (triple CNA) were much higher in frequency in TCGA (70.5%) than KNBTG (24.3%). In addition, triple CNA proved to be a common poor prognostic factor between KNBTG and TCGA. In patients who received standard treatment, median overall survival was 19.3 months for the KNBTG cohort and 15.6 months for the TCGA cohort, and this survival difference was highly associated with differences in the triple CNA frequency.
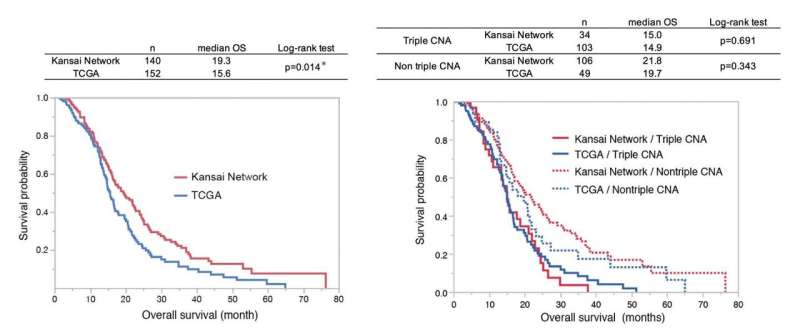
From these results, they found that triple CNA can be used as a prognosis biomarker and that the discrepancies in the distribution of triple CNA in these two cohorts could provide a full account of the cohort disparities of survival duration.
Toru Umehara from Osaka University said, "Besides GBM, several cancers in which the disproportion of the genetic landscape potentially varies the morbidity and clinical outcome have been reported. Even if the clinical outcome and genetic landscape of a disease are available from previous studies, it's important to create their own report and build an original database by region. Our research results will provide new insight into the interpretation and comparison of interregional clinical trials for GBM."
More information: Toru Umehara et al, Distribution differences in prognostic copy number alteration profiles in IDH-wild-type glioblastoma cause survival discrepancies across cohorts, Acta Neuropathologica Communications (2019). DOI: 10.1186/s40478-019-0749-8