Novel paradigm in drug development

Traditional medicines mostly function as inhibitors, attacking the disease-relevant proteins that cause cancer, by binding to their accessible pockets. Following this strategy, only ~20% of all proteins are chemically addressable, leaving some of the most relevant targets inaccessible to therapeutic development.
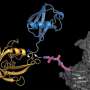
Targeted protein degradation (TPD) is a novel approach in drug development that could overcome this limitation, and currently represents a promising therapeutic strategy towards, for example, cancer treatment. TPD is based on small-molecules, generally called "degraders," which induce the degradation of proteins by re-directing ubiquitin E3 ligases towards the protein we aim to eliminate. In other words, utilizing the cell's Ubiquitin Proteasome System (UPS), which is our body's natural way of seeking out and destroying damaged proteins.
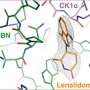
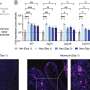
Until now TPD had been mostly studied from a structural perspective. Georg Winter's laboratory at CeMM focused on identifying and mechanistically understanding genetic determinants of sensitivity to small-molecule degraders. "We selected a representative set of five degraders, which hijack different ubiquitin E3 ligases to degrade proteins of clinical relevance, such as BRD4, CDK9, or GSPT1. Conducting resistance screens, we were able to identify genes that determine the efficacy of targeted protein degradation," explains Cristina Mayor-Ruiz, CeMM postdoc and co-first author of the study.
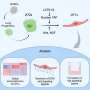
The data obtained identify central UPS regulators as essential for degrader efficacy. "When those proteins are perturbed, ubiquitin E3 ligases lose their ability to flexibly assemble and disassemble in response to cellular needs. Instead, they start tagging themselves for destruction in a process called auto-degradation. As a consequence, the tested degrader drugs fail to destabilize their target proteins and are ineffective in blocking cancer cell growth," elaborates Martin Jaeger, CeMM Ph.D. student and second co-first author of the study.
The research conducted by Cristina Mayor-Ruiz, Martin Jaeger et al. combining functional genomics and quantitative proteomics is the first study that comprehensively dissects cellular determinants of mechanistically different small-molecule degraders, bringing new light into their rational design.
"Now that degraders are entering the clinic, understanding potential resistance mechanisms may inform on ways to overcome it. The modulator gene-networks that we have identified can serve as biomarkers to support patient stratification, but also teach us a lot about fundamental aspects of the regulation and dynamics of the protein degradation machinery," says Georg Winter, CeMM Principal Investigator.
More information: Cristina Mayor-Ruiz et al, Plasticity of the Cullin-RING Ligase Repertoire Shapes Sensitivity to Ligand-Induced Protein Degradation, Molecular Cell (2019). DOI: 10.1016/j.molcel.2019.07.013