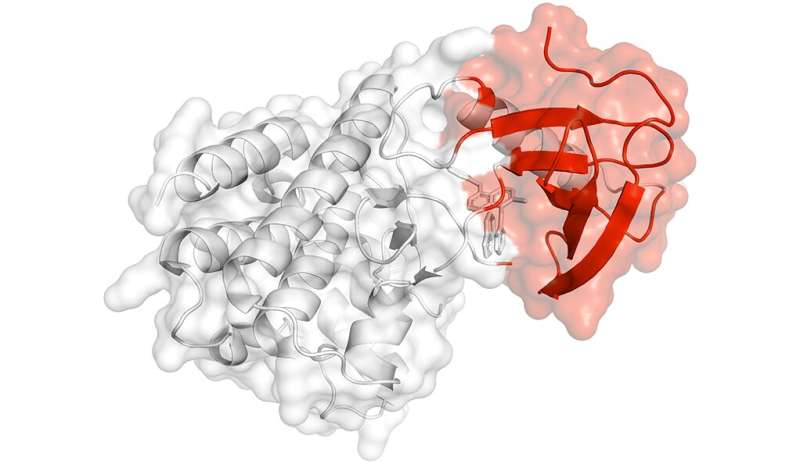
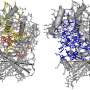
Yale researchers exploring the structure of proteins have found evidence of how some genes drive the progression of cancer. Credit: Yale University
Scientists have long searched for "driver" genes that fuel the progression of cancer, but existing technology has had a hard time separating mutations that are true drivers from others that are simply "passengers," not directly involved in spread of tumors. However, a team of Yale researchers has developed a model that marries biophysics, advanced gene sequencing technology, and statistics to identify at least 200 new genes which may drive cancer progression, they report Sept. 2 in the journal Proceedings of the National Academy of Sciences.
Most drivers identified thus far have come by identifying the over-representation of mutations in areas of genomes of cancer patients. Scientists have also studied the static structure of proteins influenced by these mutations to identify potential driver genes.
Sushant Kumar and Declan Clarke in the lab of Mark Gerstein, the Albert L Williams Professor of Biomedical Informatics and professor of molecular biophysics & biochemistry, and of computer science, went a step further.
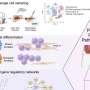
By studying dynamic 3-D models of proteins in motion, they identified key parts of proteins that harbor a significantly high frequency of cancer-associated mutations.
Using this method, they identified 434 potential driver genes, many of which have been known to researchers.
More information: Sushant Kumar et al. Leveraging protein dynamics to identify cancer mutational hotspots using 3D structures, Proceedings of the National Academy of Sciences (2019). DOI: 10.1073/pnas.1901156116
Journal information: Proceedings of the National Academy of Sciences
Provided by Yale University