Whole genome sequencing for prenatal diagnosis

The Department of Obstetrics and Gynaecology of the Faculty of Medicine at The Chinese University of Hong Kong (CUHK) has successfully introduced a new genome sequencing technique for prenatal invasive genetic diagnosis. It offers enhanced sensitivity and accuracy of diagnosing lethal and severe congenital disorders through precise detection of pathogenic microdeletion or microduplication in the fetus, compared with the current practice karyotyping analysis and chromosomal microarray analysis (as known as fetal DNA chip testing). The team conducted a study on the innovation and the findings were recently published on the journal Genetics in Medicine.
Genetic test provides more precise analysis and detects much smaller genetic abnormalities during pregnancies
Pregnant women who have experienced miscarriage or stillbirth, or those with fetal abnormalities revealed by ultrasound examination or a positive result of Down's syndrome screening, are recommended to receive further invasive prenatal diagnosis of genetic disorders in fetus.
The conventional fetal karyotyping studies the structure of the chromosomes. About two-thirds of these fetuses will have a normal karyotype result, yet at least 10 percent of them will have structural anomalies, which is known to be associated with a wide array of syndromic disorders and genetic diseases.
Professor Tak Yeung LEUNG, Chairman of the Department of Obstetrics and Gynaecology, Faculty of Medicine at CUHK explained, "If a clinically significant genetic imbalance is diagnosed in the fetus, it will allow the mother and the family to make an informed decision and assist the management of this pregnancy and those in the future. Making a specific diagnosis of these syndromic and genetic diseases prenatally is difficult, as many of them are linked to submicroscopic chromosomal abnormalities that are undetectable by conventional karyotyping."
In 2009, CUHK became the first in Hong Kong and Asia Pacific region to introduce the fetal DNA chip testing, using chromosomal microarray analysis to diagnose more than 100 common micro-deletion or duplication syndromes which would not be detected by karyotyping. This year, the Obstetrics and Gynaecology team is introducing the whole genome sequencing approach to provide an even more comprehensive and precise detection of micro-deletions or duplications.
Whole genome sequencing identifies additional and clinically relevant genomic and chromosomal abnormalities compared with fetal DNA chip testing

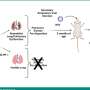
Fetal DNA chip testing and whole genome sequencing (WGS) detect genetic imbalance in a fetus or an individual by examining copy number variants (CNVs) in the chromosomes, using chorionic villus sampling cells, amniotic fluids or fetal cord blood. CNVs are the number of copies of a particular gene or sections of the genome repeated or missed which makes for variation from one individual to the next. This contributes to the distinctiveness between individuals and explanation of some human diseases.
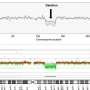
Professor Richard Kwong Wai CHOY, Associate Professor of the Department of Obstetrics and Gynaecology, Faculty of Medicine at CUHK remarked, "Fetal DNA chip testing allows quick and thorough analysis of all chromosomes in a single test. However, the CNV analysis by fetal DNA chip testing is highly reliant on the probe density of the target region. WGS is a state-of-the-art next-generation sequencing approach providing genome-wide sequence information. However, WGS with high read-depth is costly, while it also detects a large number of variants of uncertain significance. In the past 5 years, our team has developed and optimized genome-wide CNV analysis by utilizing low-pass WGS with an in-house analytic pipeline named FetalSeq and it has demonstrated robustness in detection of CNVs at higher resolution and provides additional and cytogenetically relevant information for prenatal diagnosis compared to fetal DNA chip testing. The throughput is as high as 48 samples per run."
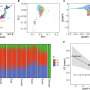
To evaluate the accuracy, efficacy, and incremental yield of FetalSeq compared with fetal DNA chip testing, the CUHK team conducted a prospective back-to-back comparison in 1,023 prenatal women referred for invasive genetic testing. The subjects enrolled in the study between late 2016 and early 2019 through two prenatal diagnosis centers under CUHK and Jinan University.
The result demonstrates that FetalSeq is able to detect all genetic aberrations detectable by fetal DNA chip testing and, furthermore, provides additional diagnostic information in 1.7 percent (17/1,023) of cases. A follow-up study confirmed the pathogenicity of those additional CNVs detected by FetalSeq.
Dr. Elvis Zirui DONG, Research Assistant Professor of the Department of Obstetrics and Gynaecology of the Faculty of Medicine at CUHK remarked, "With genome-wide resolution, FetalSeq shows the potential advantage in identifying genetic etiologies in human diseases attributed to chromosomal disorders, pathogenic CNVs or monogenic disorders such as Duchenne muscular dystrophy. The technology provides more precise prognostic information to the pregnant women and their families for the further management."
CUHK Obstetrics and Gynaecology team pioneers in prenatal genetic diagnosis
The Department of Obstetrics and Gynaecology of the Faculty of Medicine at CUHK is one of the leaders in prenatal genetic diagnosis in Hong Kong. Since the introduction of fetal DNA chip testing for identification of numerical disorder (i.e., Down syndrome) and microdeletion or duplication syndrome in 2009, the Department has been servicing a substantially increasing demand for invasive genetic testing. In June 2019, the Hospital Authority began to offer fetal DNA chip testing to all Hong Kong pregnant women with high-risk pregnancies free of charge.
A recent CUHK study on the spectrum of clinically significant CNVs in prenatal diagnosis by fetal DNA chip testing provided evidence that pathogenic genetic aberrations occur 1 in 64 pregnancies undergoing invasive testing.