Researchers perform thousands of mutations to understand amyotrophic lateral sclerosis

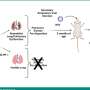
Amyotrophic lateral sclerosis (ALS) is a devastating and incurable nervous system disease that affects nerve cells in the brain and spinal cord, causing loss of muscle control and normally death within a few years of diagnosis. In ALS, as in other neurodegenerative diseases, specific protein aggregates have long been recognized as pathological hallmarks, but it is not clear whether they represent the actual cause of the disease. Indeed, alleviating aggregation has repeatedly failed as a therapeutic strategy for treating neurodegenerative diseases such as Alzheimer's.
In order to cast more light on this issue, researchers at the Center of Genomic Regulation (CRG) and the Institute of Bioengineering of Catalonia (IBEC) have applied a novel approach called deep mutagenesis with unexpected results. "By studying all possible mutations in a protein, we have a much more reliable way to understand toxicity, and we are excited to move on to many more proteins implicated in neurodegenerative diseases," says Benedetta Bolognesi, IBEC researcher and first author of the paper.
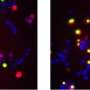
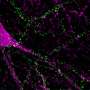
In a collaboration between the labs of ICREA research professors Ben Lehner and Gian Tartaglia, Benedetta Bolognesi and Andre J. Faure focused on TDP-43, a protein that aggregates in the motor neurons of nearly all ALS patients. They made over 50,000 mutants of TDP-43 and tracked their toxicity to yeast cells. Researchers found that mutant forms that aggregated were actually less toxic than other versions of the protein which instead were forming unusual liquid species in the cells. "This is the exact opposite of what we expected," Lehner says, and challenges a lot of the assumptions in this field.
It still remains to be established whether aggregation of TDP-43 is also protective in mammalian cells and neurons, something Bolognesi is working on, but if this proves to be the case, it means we will have to entirely change the way we therapeutically address ALS if we want to have significant effects.
More information: Bolognesi, B., et al (2019). The mutational landscape of a prion-like domain. Nature Communications 10, 4162