KAT6A syndrome: advances on the genetic bases and clinical picture of a rare disease

A research team has described five new cases of a rare disease known as KAT6A syndrome, of which there are only 80 dominant cases worldwide. This neurological and developmental disorder, caused by alterations in the lysine acetyltransferase 6A gene (KAT6A), involves intellectual disability, language impairment, low muscle tone, cardiovascular malformation and eye defects, among other symptoms.
The study, published in the journal Orphanet Journal of Rare Diseases, is co-led by researchers Daniel Grinberg and Susanna Balcells and Roser Urreizti from the Faculty of Biology of the University of Barcelona and the Rare Diseases Networking Biomedical Research Centre (CIBERER), and the experts Estrella López-Martín and Eva Bermejo-Sánchez, from the Institute of Rare Diseases Research (IIER) from Carlos III Health Institute (ISCIII).
The research team confirmed in five new patients some of the features of the clinical picture related to the pathology, apart from describing new symptoms and affectations in the development of the disease. In particular, the authors observed new symptoms such as cryptorchidism (testicular atrophy), syndactyly (fused fingers) and trigonocephaly (deformation in the frontal part of the cranium), apart from the emergence of recurrent infections.
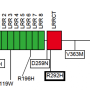
The researchers identified the role of genetic mutations in this minority disease. Though the syndrome was first associated with truncating mutations, the new study describes a variant of the disease without cardiac symptoms that is related to missense mutations, in which the change of a single base pair causes the substitution of a different amino acid in the resulting protein.
Four out of the five patients described in this study showed missense substitutions, and in the fifth case, they observed a missense mutation that did not follow cardiac symptoms.
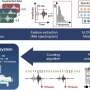
In silico studies and informatic simulations confirmed that the missense mutation generated a partial process of splicing that caused the premature truncating of the protein that gave origin to the pathology.
The new study will contribute to better understanding the natural history of this syndrome and expanding the clinical phenotype of patients, advances that are essential to delimit the phenotypical characterization and definition of this rare disease. According to the authors, these findings will help to increase the chances to diagnose other affected patients earlier.
More information: Roser Urreizti et al. Five new cases of syndromic intellectual disability due to KAT6A mutations: widening the molecular and clinical spectrum, Orphanet Journal of Rare Diseases (2020). DOI: 10.1186/s13023-020-1317-9