Speedy antibiotic susceptibility tests for high-priority pathogens
At the core of the antibiotic-resistance crisis is the lack of a rapid and general antibiotic susceptibility test (AST) that can assess the infecting pathogen's sensitivity to antibiotics and thus inform treatment decisions directly at the point of care. In two new papers publishing March 19 in the open-access journal PLoS Biology, researchers from the lab of Rustem Ismagilov at the California Institute of Technology have developed two distinct, highly innovative AST methods for directly measuring a pathogen's susceptibility to beta-lactams (a class of antibiotics that includes penicillin) on extremely short time scales (about 30 minutes).
Rapid "genotypic" tests detect the presence of antibiotic-resistance genes in a pathogen, but these tests can detect only a small handful of the myriad potential antibiotic-resistance mechanisms. By contrast, "phenotypic" tests directly measure a pathogen's response to an antibiotic and thus are the gold standard. However, most phenotypic tests require the pathogen to be cultured and are thus too slow (taking hours to days).
New phenotypic tests quantify small changes in the concentration of the pathogen's nucleic acids after antibiotic exposure, and are promising for fast-growing pathogens and antibiotics that affect DNA replication. However, there has not been a path for developing these rapid ASTs for slow-growing pathogens or antibiotics that do not immediately impact DNA replication, such as beta-lactams (one of the most important classes).
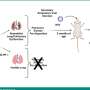
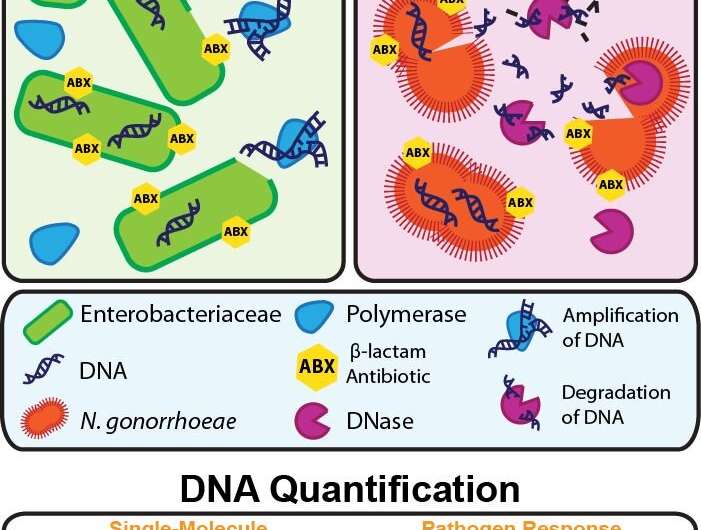
This breakthrough in AST development was made possible by measuring the accessibility of a bacterium's DNA after exposure to antibiotics. Beta-lactam antibiotics work by inhibiting a bacterium's ability to make and repair its cell wall. The researchers therefore reasoned that the bacterium's resistance or susceptibility to beta-lactams can be assessed by measuring how much of the DNA is no longer protected by an intact cell wall thus directly linking—for the first time ever—cell wall damage to a rapid DNA readout.
"One of the most exciting parts of this project was realizing that we could use a DNA readout to measure antibiotic susceptibility even if the antibiotic being tested did not directly impact the bacterium's DNA replication" says Nathan Schoepp, a former Ph.D. student in the Ismagilov Lab and a co-author on both studies.
Each AST method represents an innovative approach needed to overcome the inherent challenges of rapidly determining a pathogen's susceptibility to beta-lactams. Each paper includes pilot experiments with patient samples, demonstrating the relevance and potential of these tests to eventually be translated to clinical settings.
Importantly, the two methods target two of the three top priorities on the CDC's list of greatest antimicrobial resistance threats: carbapenem-resistant Enterobacteriaceae (CRE) and Neisseria gonorrhoeae.
"At the end of the day we aspire to design diagnostic tests that are useful in a clinical context," says Emily Savela, a Caltech Ph.D. student and a co-author on both studies. "There is a lot of complexity added when working directly with patient samples and point-of-care timescales. Our research team works to identify and incorporate these added variables in the early stages of development."
The clinical work in these two papers was made possible through the Caltech team's collaborations with co-authors Dr. Jeffrey Klausner (AIDS Healthcare Foundation and UCLA), Dr. Olusegun O. Soge (Global Health STI Laboratory, University of Washington), and Dr. Omai Garner (UCLA).
Getting these new AST methods into the clinic will still require a huge amount of work; however, the research team is optimistic that these new capabilities finally lay the pathway toward a general, rapid test of antibiotic susceptibility.
More information: Savela ES, Schoepp NG, Cooper MM, Rolando JC, Klausner JD, Soge OO, et al. (2020) Surfactant-enhanced DNA accessibility to nuclease accelerates phenotypic β-lactam antibiotic susceptibility testing of Neisseria gonorrhoeae. PLoS Biol 18(3): e3000651. doi.org/10.1371/journal.pbio.3000651
Schoepp NG, Liaw EJ, Winnett A, Savela ES, Garner OB, Ismagilov RF (2020) Differential DNA accessibility to polymerase enables 30-minute phenotypic β-lactam antibiotic susceptibility testing of carbapenem-resistant Enterobacteriaceae. PLoS Biol 18(3): e3000652. doi.org/10.1371/journal.pbio.3000652