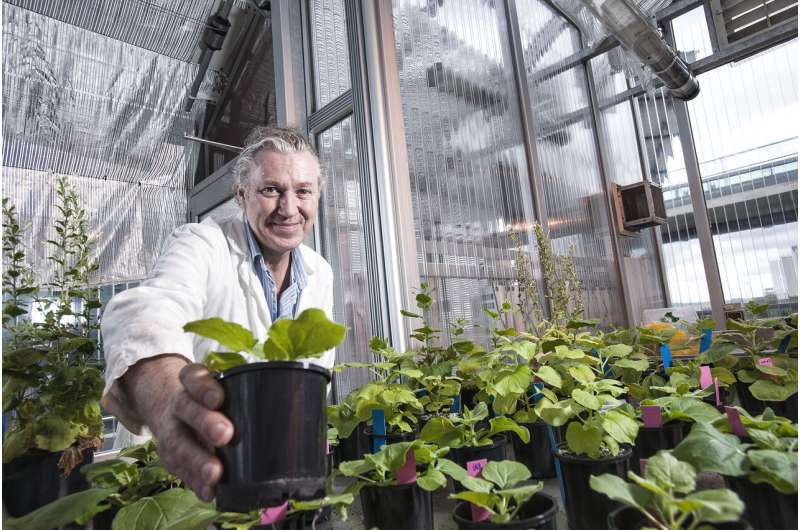
Professor Peter Waterhouse. Credit: QUT
Professor Peter Waterhouse, an Australian Laureate Fellow, has developed a roadmap for biologists and biotechnologists who are turning to the new technique of producing antibodies, vaccines and therapeutics, including those for COVID-19, in plants.
Professor Waterhouse, from Laboratory of Molecular Genetics (LMG) at QUT, said an international biotech company had already been given fast-tracked access to the entire chromosome-level genome sequence of the Australian-native plant Nicotiana benthamiana.
"We are now making this unpublished information available to any team working on fighting the COVID-19 pandemic," Professor Waterhouse said
"This will allow them to 'tweak' the genome to produce better quality vaccines and therapeutics that are tailor-made for use in humans."
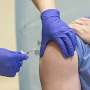
COVID-19 has caused the largest and most deadly pandemic since the 1918 Spanish flu and a large number of public and private research groups are working around the clock to develop protein-based diagnostic products and vaccines to fight the pandemic.
One major problem, once these are developed, is their production in bulk and at a low cost.
One solution is to use plants as biofactories for the production. Plants can be grown in large amounts using simple agricultural technologies, that are within reach of developing countries that lack sophisticated protein production methods, such as those using animal cell cultures. This discipline is known as "molecular farming."
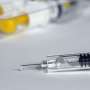
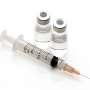
Biotechnologists can use the plant by injecting or infiltrating the DNA instructions of how to make the antibody or vaccine into its leaves and the plant does the rest—producing the vaccine in its cells and sap.
The plant, Nicotiana benthamiana, is being used all over the world as the vaccine plant biofactory of choice and the genome sequencing has been led by Professor Waterhouse in partnership with the European Horizon2020 Newcotiana consortium.
"We started the project with the Newcotiana consortium three years ago with the idea of making plants able to make larger quantities and better qualities of the vaccine and antibodies when COVID-19 was not known," Professor Waterhouse said.
"As we go forward, there will always be a need to rapidly respond to new strains of viruses as they emerge. In recent years we have seen SARS, MERS, and now COVID-19.
"It's really fortunate that we are reaching the level of understanding of this biofactory plant and have progressed the Newcotiana H2020 project as far as we have, enabling and providing new and improved ways to meet current and future challenges."
Professor Waterhouse's work, which can be used in a wide range of projects from boosting plant disease resistance, to creating better vaccines and antibodies, has been highlighted in a new series of Australian Academy of Science.
The plant, known in the lab as "benth," is an ancient native tobacco plant that naturally grows only in Northern and Central Australia, and may also hold the key to growing crops that can withstand disease and climate change.
It was first described by the ship's surgeon aboard HMS Beagle (of Charles Darwin fame) on its third voyage around Australia and in 1936 Sir John Cleland collected the seed, from a plant growing in Central Australia, which has been propagated and passed from lab to lab throughout the world.
The benth genome has about 60,000 genes, about twice the number of an ordinary plant.
"It's a special plant because it is being used for a wide spectrum of vaccines and antibodies, including those for Ebola and now for COVID-19," Professor Waterhouse said. "Plants don't usually produce antibodies, it's something that animals do."
Professor Waterhouse, from the Centre for Agriculture and the Bioeconomy and the School of Biology and Environmental Science, has been collecting and studying different ecotypes of this plant and made an expedition to the centre of Australia to find the plant. His team is also using benth to engineer disease-resistant crops.
The N. benthamiana genome sequence can be accessed at https://nbenth.com/annotator/index.
More information: More information on the Newcotiana project can be found on newcotiana.org
Provided by Queensland University of Technology