Inhibition of sphingolipid metabolism and neurodegenerative diseases
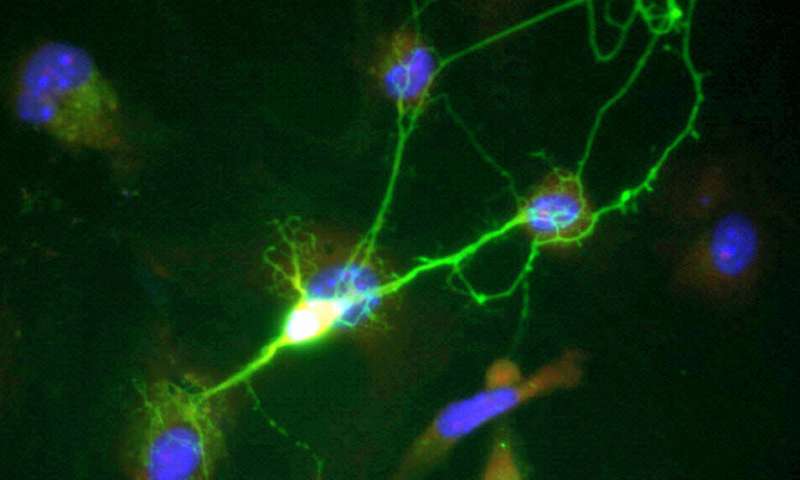
Disrupting the production of a class of lipids known as sphingolipids in neurons improved symptoms of neurodegeneration and increased survival in a mouse model, according to new research led by the joint laboratory of Robert Farese, Jr. and Tobias Walther at Harvard T.H. Chan School of Public Health and Howard Hughes Medical Institute.
The findings, published online April 13, 2020 in the Proceedings of the National Academy of Sciences (PNAS), could help in the development of therapies for a range of neurodegenerative diseases.
"This work began in yeast, and serendipitously we discovered that mutations related to neurodegeneration in humans led to abnormalities in cell lipid metabolism. Most investigators don't look at the lipids, and we were quite excited and surprised," said Walther, professor of molecular metabolism and executive director of the Harvard Chan Advanced Multi-omics Platform.
In the study, the Farese & Walther Laboratory identified a link between sphingolipid metabolism and a mutation that affected vesicle trafficking, the process by which molecules are transported to different parts of a cell. Defects in the trafficking process are known to play a role in neurodegenerative diseases, but the exact mechanism of the effect is not understood.
The Farese & Walther Laboratory has previously focused their research efforts on the Golgi-associated retrograde protein (GARP) complex. Earlier studies in yeast and flies showed that mutations in GARP proteins lead to sphingolipid abnormalities and stunted cell growth, and that these effects could be reversed by inhibiting sphingolipid metabolism. Many neurological diseases are caused by mutations in genes that are involved in lipid metabolism, and changes in lipid metabolism have been reported in amyotrophic lateral sclerosis (ALS), Parkinson's disease, and Alzheimer's disease. With that in mind, the Farese & Walther Laboratory investigated whether modulating sphingolipid metabolism affects cell dysfunction as it relates to neurodegenerative diseases.
The research team, which included Harvard Chan scientists Constance Petit and Jane Lee, used a model known as wobbler mice, which have a mutation in a specific GARP protein called VSP54. That mutation causes motor-neuron disease similar to ALS. They found that sphingolipid molecules that were toxic to cells accumulated in spinal cords of wobbler mice, as well as in embryonic fibroblasts, a type of cell that was cultured from the mice. Furthermore, they showed that certain molecules involved in sphingolipid metabolism were misplaced in neurons from wobbler mice.
The team then found that treating the mice with the sphingolipid synthesis inhibitor myriocin, which is already used as an antifungal and immunosuppressive drug, prevented the buildup of sphingolipids, reduced their toxic effects, improved wellness scores in the mice, and ultimately extended the animals' lifespan.
The results indicate that compromised sphingolipid metabolism in GARP mutations are a potential cause of neurodegeneration and that correcting defects in sphingolipid metabolism might restore neuronal function. Sphingolipid metabolism may therefore be an important target for therapeutic development for neurological disorders associated with mutations in membrane trafficking.
"We are cautiously optimistic," Farese said. "Perhaps targeting lipid abnormalities provides a new therapeutic angle for some diseases of neurodegeneration."
More information: Constance S. Petit el al., "Inhibition of sphingolipid synthesis improves outcomes and survival in GARP mutant wobbler mice, a model of motor neuron degeneration," PNAS (2020). www.pnas.org/cgi/doi/10.1073/pnas.1913956117