Global spread of the multi-resistant pathogen Stenotrophomonas maltophilia
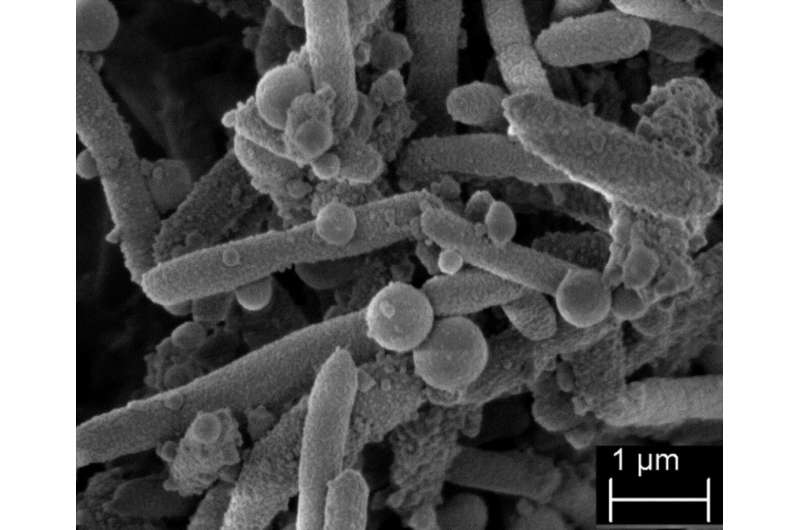

An international consortium found a remarkable global spread of strains of a multi-resistant bacterium that can cause severe infections—Stenotrophomonas maltophilia. The study, published under the supervision of the Research Center Borstel Leibniz Lung Center (FZB), provides for the first time a systematic understanding of the global phylogeny of S. maltophilia strains and shows ways to efficiently monitor the pathogen using a genomic classification system. DZIF scientists from Lübeck, Borstel and Braunschweig are involved in the study.
S. maltophilia strains occur in several natural and human associated ecosystems. The bacterium was long regarded as relatively unproblematic but is now considered to be one of the most feared hospital pathogens, as it frequently causes infections and is resistant to a number of antibiotics. This can be particularly dangerous for immune-compromised patients or for patients with underlying inflammatory lung diseases such as cystic fibrosis. Although almost any organ can be affected, infections of the respiratory tract, bacteraemia or catheter-related infections of the bloodstream are the most common. In view of the increasing importance of this pathogen and the often-severe clinical consequences of an infection, knowledge about the virulence factors and about the local and global transmission of S. maltophilia bacteria is urgently needed.
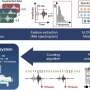
Scientists from a total of eight countries initially established a genotyping method that enables the standardised analysis of the different genomes of S. maltophilia strains. The DZIF teams around Prof. Stefan Niemann (FZB), Prof. Jan Rupp, (Clinic of Infectiology and Microbiology, Campus Lübeck) and Prof. Ulrich Nübel from the Leibniz Institute DSMZ (German Collection of Microorganisms and Cell Cultures GmbH ) in Braunschweig were involved.
The scientists found that the S. maltophilia complex can be divided into a total of 23 lineages with different prevalence levels. One particular line of descent appeared worldwide and had the highest rate of human-associated strains. This "Sm6" strain was also characterised by the presence of key virulence genes and resistance genes. "This suggests that a specific gene configuration may promote the spread of different S. maltophilia subtypes in the hospital setting, i.e. under antimicrobial treatment," says Matthias Gröschel, lead author of the study.
Transmission analysis also identified several potential outbreak events of genetically closely related strains that were isolated within days or weeks in the same hospitals. "Combined with studies on other pathogens, our results show how systematic genome-based monitoring of S. maltophilia and other pathogens in hospital settings can help detect transmission pathways and improve infection control," Thomas Kohl, senior author of the study at FZ Borstel, adds.
More information: Matthias I. Gröschel et al, The phylogenetic landscape and nosocomial spread of the multidrug-resistant opportunist Stenotrophomonas maltophilia, Nature Communications (2020). DOI: 10.1038/s41467-020-15123-0