Signatures in blood reveal how sepsis patients will respond to condition
Scientists have identified molecular signatures in the immune component of the blood which indicate how patients in intensive care with sepsis, septic shock and systemic inflammatory response syndrome are likely to respond to the conditions.
The work—led by Public Health England and involving Cardiff and Nottingham Trent universities—means that for the first time clinicians would be able to test and pre-emptively manage and treat patients based on their immune profiles.
It is hoped the study could also pave the way for new therapies, as the researchers were able to identify key molecules in the immune system which could become new drug targets.
They also believe this approach could be applied to COVID-19, given it manifests as a sepsis-like disease in the more severe cases.
Dr. Tamas Szakmany, senior lecturer in intensive care at Cardiff University, said: "Sepsis on the intensive care unit can present in several ways and we have learnt that defining the group of patients based on solely clinical parameters is difficult.
"Detailed understanding of the molecular response to infection will help us to treat those patients with novel therapies, who are most likely to benefit from these experimental approaches.
"We believe our bioinformatics model can be readily applied when looking for new biomarkers which could help us to determine the severity and prognosis of the new COVID-19 disease. Stratification of patients into risk groups would enhance our abilities to direct novel treatments to those who most benefit from them."
Sepsis is a life-threatening condition—requiring admission to intensive care—and occurs when the immune system overreacts to an infection and starts to damage the body's tissues and organs.
It is a major healthcare problem in the UK and accounts for a quarter of intensive care admissions in the UK.
Despite this there is a lack of knowledge of the immune processes involved in sepsis or the clinically relevant molecules at play. This would help clinicians to distinguish between sepsis and systemic inflammatory response syndrome SIRS—which are very similar—to improve patient management through more appropriate treatment as well as help to identify potential new therapies.
The team analysed molecules in the white blood cells, which function as part of the immune system, of patients with sepsis, septic shock—the most severe form of sepsis—and SIRS.
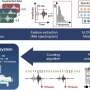
Using Nottingham Trent University-led machine learning and artificial intelligence, they were then able to develop molecular signatures that were able to predict the outcome of patients based on their immune response, the condition they had and whether it had started in the lungs or the abdomen.
Work is ongoing at PHE and Cardiff to further develop these signatures into clinically-useful diagnostic tools.
Nottingham's Professor Graham Ball said: "This approach provides new insights into how patients respond to these serious conditions based on their immune response and the molecular processes that define and drive disease progression. Our work highlights the importance of examining these molecular immune responses in determining outcome for patients.
"Another important aspect of this study is that the molecular processes we identified are similar to those defining patient outcome in COVID-19. As such our methods could potentially be used to predict response and outcome for these patients too."
More information: Dong Ling Tong et al. Development of a Bioinformatics Framework for Identification and Validation of Genomic Biomarkers and Key Immunopathology Processes and Controllers in Infectious and Non-infectious Severe Inflammatory Response Syndrome, Frontiers in Immunology (2020). DOI: 10.3389/fimmu.2020.00380