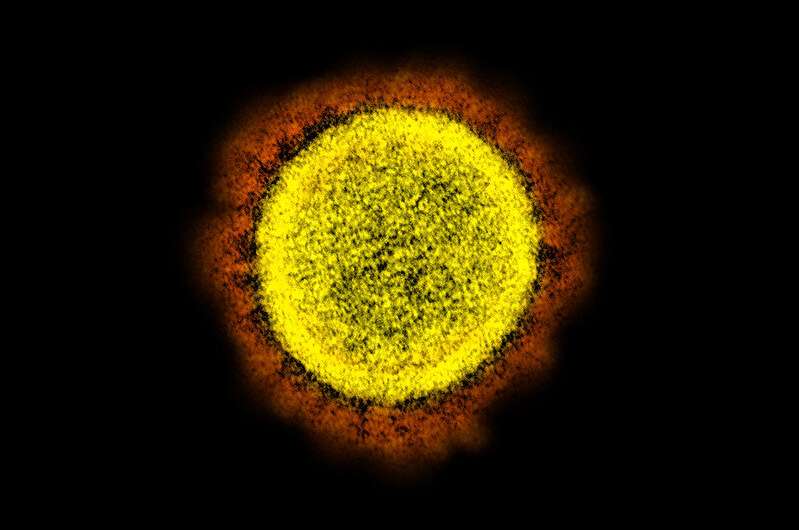
Novel Coronavirus SARS-CoV-2 Transmission electron micrograph of SARS-CoV-2 virus particles, isolated from a patient. Image captured and color-enhanced at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Credit: National Institute of Allergy and Infectious Diseases, NIH
Scientists have shown that two species of seasonal human coronavirus related to SARS-CoV-2 can evolve in certain proteins to escape recognition by the immune system, according to a study published today in eLife.
The findings suggest that, if SARS-CoV-2 evolves in the same way, current vaccines against the virus may become outdated, requiring new ones to be made to match future strains.
When a person is infected by a virus or vaccinated against it, immune cells in their body will produce antibodies that can recognize and bind to unique proteins on the virus' surface known as antigens. The immune system relies on being able to 'remember' the antigens that relate to a specific virus in order to provide immunity against it. However, in some viruses, such as the seasonal flu, those antigens are likely to change and evolve in a process called antigenic drift, meaning the immune system may no longer respond to reinfection.
"Some coronaviruses are known to reinfect humans but it is not clear to what extent this is due to our immune memory fading or antigenic drift," says first author Kathryn Kistler, a Ph.D. student at the Vaccine and Infectious Disease Division, Fred Hutchinson Cancer Research Center, Seattle, US. "We wanted to investigate whether there is any evidence of coronaviruses related to SARS-CoV-2 evolving to evade our immune responses."
The research team looked at the four seasonal human coronaviruses (HCoVs) which are related to SARS-CoV-2 but typically cause milder symptoms, such as the common cold. HCoVs have been circulating in the human population for 20-60 years, meaning their antigens would likely have faced pressure to evolve against our immune system.
The team used a variety of computational methods to compare the genetic sequences of many different strains of the viruses, enabling them to see how they have evolved over the years. They were especially interested in any changes that might have occurred in viral proteins that could contain antigens, such as spike proteins—the ray-like projections on the surface of coronaviruses that are particularly exposed to our immune system.
The researchers found a high rate of evolution in the spike proteins of two of the four viruses, OC43 and 229E. Nearly all of the beneficial mutations appeared in a specific region of the spike proteins called S1, which helps the virus infect human cells. This suggests that reinfection by these two viruses can occur as a result of antigenic drift as they evolve to escape recognition by the immune system.
The team also estimated that beneficial mutations in the spike proteins of OC43 and 229E appear roughly once every two to three years, about half to one-third of the rate seen in the flu virus strain, H3N2.
"Due to the high complexity and diversity of HCoVs, it is not entirely clear if this means that other coronaviruses, such as SARS-CoV-2, will evolve in the same way," explains senior author Trevor Bedford, Principal Investigator and Associate Member at the Vaccine and Infectious Disease Division, Fred Hutchinson Cancer Research Center. "The current vaccines against COVID-19, while highly effective, may need to be reformulated to match new strains, making it vital to continually monitor the evolution of the virus' antigens."
More information: Kathryn E Kistler et al, Evidence for adaptive evolution in the receptor-binding domain of seasonal coronaviruses OC43 and 229E, eLife (2021). DOI: 10.7554/eLife.64509
Journal information: eLife
Provided by eLife