Curcumin for amyloidosis and lipid metabolism—a novel insight
Curcumin is a polyphenol compound produced by plants of the Curcuma longa species and has been reported to have many physiological activities, which include anti-oxidation, anti-inflammatory, anti-cancer, and anti-amyloid properties. However, the mechanism and network of action are not completely clear. Amyloidosis is a group of diseases characterized by abnormal aggregates of proteins, known as amyloid fibrils, and subsequent deposition in various tissues and organs, such as Alzheimer's disease, immunoglobulin light chain amyloidosis.
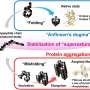
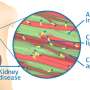
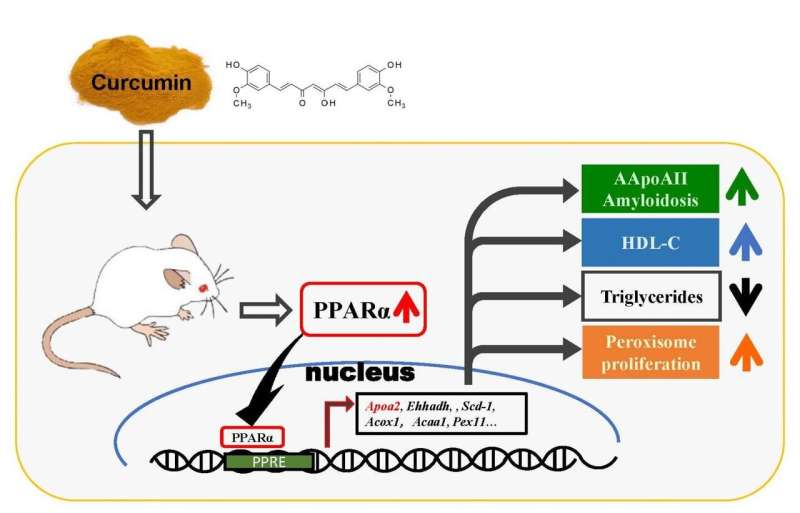
In previous studies, curcumin has been shown to suppress the aggregation and cytotoxicity of many amyloid proteins in vitro, such as amyloid ß (Aß), α-synuclein, transthyretin, and prion protein, and has also been reported to inhibit the deposition of Aß fibrils in a mouse model of Alzheimer's disease. The group investigated amyloid deposition and molecular changes in a mouse model of amyloid apolipoprotein A-II (AApoAII) amyloidosis, in which mice were fed a curcumin-supplemented diet. In this research, it was found that curcumin intake elevated ApoA-II and HDL-cholesterol concentration in plasma by activating the peroxisome proliferator-activated receptor alpha (PPARα) signaling pathway, resulting in increased AApoAII amyloid deposition and peroxisome proliferation. These findings demonstrate the novel agonistic effect of curcumin on PPARα, which is an important transcription factor for lipid metabolism, and may have far-reaching significance for the treatment of amyloidosis and other metabolic disorders.
It was reported that high-fat diet supplement aggravates a variety of amyloid deposition including Aß in Alzheimer's disease model mice, but a link between lipid metabolism and the development of amyloidosis has not been completely established. These results provide a promising molecular target to understand the molecular mechanism of amyloidogenesis, which the activation state of PPARα pathway may be a bridge to connect the change of lipid metabolism level and the degree of amyloid deposition. In addition, it has been regarded that curcumin, as an agonist of PPARγ, exerts anti-inflammatory, anti-cancer and antioxidant activities in the past. However, this study demonstrates that curcumin is a PPARα/γ dual activator and may affect expression levels of proteins involved in amyloid deposition and other metabolism functions in a complex manner. By focusing on the PPARα pathway, the group hope to provide an opportunity to reconsider the mechanism of the physiological effects of curcumin.
In the next stage, the group would like to clarify how curcumin activates the PPARα signaling pathway in vitro and confirm whether the activation of PPARα can affect amyloid deposition on other types of amyloidosis in vivo (Alzheimer's disease, ATTR amyloidosis etc.). The goal in this research is to elucidate the molecular pathways involved in the pathogenesis of amyloidosis in vivo and to develop effective therapeutic or preventive methods against the development of amyloidosis. As a future study, the group hopes to fully understand the molecular fluctuations of PPARα-activated cells and verify the effectiveness of interventions in these pathways for various metabolic diseases.
More information: Jian Dai et al, Curcumin promotes AApoAII amyloidosis and peroxisome proliferation in mice by activating the PPARα signaling pathway, eLife (2021). DOI: 10.7554/eLife.63538