Targeting a new antibody supersite key to COVID immunity
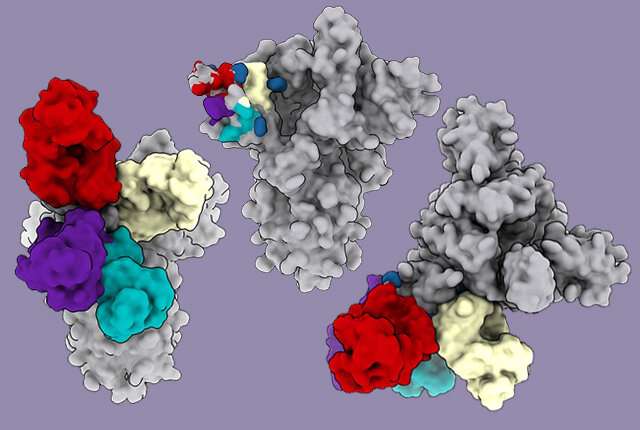
Scientists are learning that a lesser-studied region on the pandemic coronavirus is recognized by COVID-19 infection-fighting antibodies. These antibodies were identified in blood samples from previously infected patients, and were found to potently prevent the virus from infecting cells.
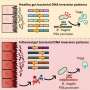
The coronavirus spike protein is the key that unlocks the door to the cell, and antibodies bind to the spike protein to jam this function. Much attention has been given to studying antibodies that target the receptor-binding domain on the coronavirus spike protein. (The receptor-binding domain of the spike is responsible for triggering the merging of the virus with a host cell to achieve a takeover.)
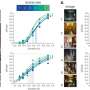
However, some of the recovered patients' antibodies blocked the coronavirus by binding to a different place on the virus spike—the N-terminal domain. These antibodies were as strong as those that bind to receptor-binding domain, a recent study shows.
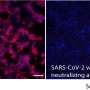
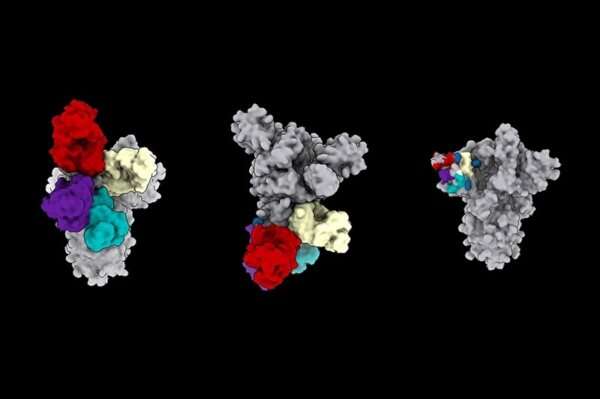
Using electron cryo-microscopy (cryoEM) to map where these antibodies bound showed that all the antibodies that prevent infection bind a single place on the N-terminal domain. The research published in Cell demonstrated that these antibodies protected Syrian hamsters from SARS-CoV-2, the coronavirus that causes COVID-19 in people.
Additional recent findings indicate that the virus is slowly defying these antibodies that people are acquiring. The virus is adapting to these antibodies by accumulating mutations that help the virus escape these defenses, becoming so-called variants-of-concern.
Some of these variants, such as those first detected in the United Kingdom and South Africa, contain mutations that appear to make the virus less vulnerable to the neutralizing power of the N-terminal domain antibodies.
"Several SARS-CoV-2 variants harbor mutations within their N-terminal domain supersite," the researchers noted. "This suggests ongoing selective pressure."
They added that investigating these neutralization escape mechanisms is revealing some unconventional ways the N-terminal domain on the virus is acquiring antibody resistance, and are why N-terminal domain variants warrant closer monitoring.
The senior authors on the Cell paper are David Veesler, associate professor of biochemistry at the University of Washington School of Medicine in Seattle, as well as Matteo Samuele Pizzuto and Davide Corti of Humabs Biomed SA, a subsidiary of Vir Biotechnology. The lead authors are Matthew McCallum of the UW medical school's Department of Biochemistry, and Anna De Marco of Humabs Biomed.
The N-terminal domain antibodies in this study were derived from memory B cells, which are white blood cells that can persistently recognize a previously encountered pathogen and re-launch an immune response.
N-terminal domain-specific antibodies likely act in concert with other antibodies to wage a multi-pronged uprising against the coronavirus. The N-terminal domain antibodies appear to inhibit virus-cell fusion. In conjunction, another part of the antibody, called a constant fragment, might also activate some of the body's other approaches to eliminating the virus.
"This study shows that NTD-directed antibodies play an important role in the immune response to SARS-CoV-2 and they appear to contribute a key selective pressure for viral evolution and the emergence of variants," said Veesler
Continuing research on the N-terminal domain neutralizing antibodies may lead to improved therapeutic and preventive anti-viral drugs for COVID-19, as well as inform the design of new vaccines or the evaluation of current ones.
For example, patients who have recovered from COVID-19 and later received a first dose of an mRNA vaccine might experience a boost in their N-terminal domain neutralizing antibodies. Also, a cocktail of antibodies that target different critical domains on the coronavirus might also be a promising approach for medical scientists to examine to see if it provides broad protection against variant strains.
The researchers stressed that, although current vaccines "are being deployed at an unprecedented pace, the timeline for large-scale manufacturing and distribution to a large enough population for community immunity still remains uncertain."
Antiviral drugs, they explain, are expected to play a role in controlling disease during the ongoing pandemic. They are likely to be particularly helpful, according to the researchers, for unvaccinated individuals and for those who didn't get a strong enough immune response from their vaccinations.
Antivirals could also prove vital when immunity from previous infection or from vaccination wanes, or as mutant strains that break through the shield of vaccination emerge.
More information: Matthew McCallum et al, N-terminal domain antigenic mapping reveals a site of vulnerability for SARS-CoV-2, Cell (2021). DOI: 10.1016/j.cell.2021.03.028