Redox imaging allows measurement of drug responses in lab-grown cancer samples
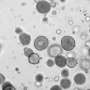
Organoids are tiny three-dimensional cellular assemblies that are grown in a laboratory from tissue-specific cells. They are particularly interesting to biologists because of their ability to mimic the characteristics of the original tissues. If scientists extract cells from a tumor, then they can grow cancer organoids that mimic the characteristics of the source tumor.
This possibility for individual-level studies of tumor properties makes cancer organoids an exciting tool from the perspective of an emerging field called precision cancer medicine. Daniel Gil of the University of Wisconsin (UW) and Morgridge Institute for Research explains, "Precision cancer medicine is this idea that we can use data to identify better treatments for each patient, and a particularly powerful approach is to use patient-derived cancer organoids, essentially 'tumors-in-a-dish,' to test drugs before giving them to a patient." Gil notes that a key prerequisite for realizing this idea is finding a way to determine whether or not the organoids are responding to a treatment. Existing methods for making this determination provide limited information about drug effects or require destroying the organoids.
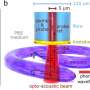
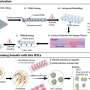
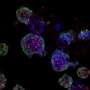
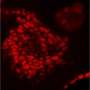
A research team including Gil, and directed by Melissa Skala of the Morgridge Institute for Research, therefore aimed to develop an informative, nondestructive way of imaging cancer organoid responses to drug treatments. The researchers chose to focus on an optical imaging technique called redox imaging, which relies on fluorescent signals from metabolites in patient-derived cancer organoids.
The researchers hoped to find a set of fluorescent signals readable with redox imaging that would provide insights into the size, shape, and function of each organoid. By analyzing how drug treatments affected such signals, they aimed to develop a screening tool for assessing organoid responses to drug treatments. Their findings are published in the Journal of Biomedical Optics.
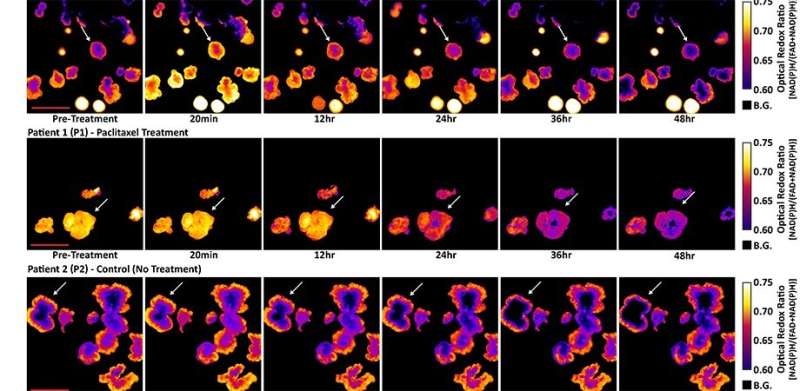
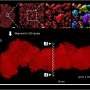
For their experiments, the researchers used redox imaging to record signals from two metabolites, called NAD(P)H and FAD, in two separate colorectal cancer organoid lines over a period of 48 hours after drug treatment. As expected, drugs known to kill cancer cells measurably altered the redox signals from the metabolites in ways that suggested a reduction in cell growth rates. The researchers noted that tracking response within a single organoid provided more sensitive information about drug responsiveness than pooled analyses across all organoids in a dish.
Further, redox imaging data analysis allowed the researchers to identify organoid subpopulations that responded to drug treatments in distinct ways. This finding aligns with what scientists know about the heterogeneity of cells within tumors. Gil explains, "Tumors aren't made up of a single cell type, but rather many cell types that often have differing drug sensitivities." He also noted that this heterogeneity can complicate the task of identifying the optimal treatment for a tumor, but the ability of patient-derived cancer organoids to capture tumoral cell heterogeneity may help scientists and clinicians learn to overcome such complications.
This study shows that redox imaging enables nondestructive measurement of drug responses in patient-derived cancer organoids. Gil notes that this approach is well-suited to evaluating large populations of organoids because it's fast and informative. "Redox imaging is a valuable tool that will allow researchers to observe the dynamic nature of drug responses over time," says Gil. Skala remarks, "This is an important step to bring this technology to the clinic in a usable, cost effective way. We hope this can guide treatment choices for more patients."
More information: Daniel A. Gil et al, Patient-derived cancer organoid tracking with wide-field one-photon redox imaging to assess treatment response, Journal of Biomedical Optics (2021). DOI: 10.1117/1.JBO.26.3.036005