New method maps brain activity despite uncertainties in patient head structure
Skoltech researchers have proposed a method for interpreting brain activity data that proved to be up to five times more accurate than the conventionally used technique in cases when MRI data contained artifacts or only a low-resolution head model was available. Reported in IEEE Transactions on Medical Imaging, the findings are of use for treating drug-resistant epilepsy and understanding cognitive processes in the healthy brain, including how it responds to visual stimuli and records new words.
Mapping brain activity is the standard way to determine which parts of the brain are involved in a specific cognitive task—for example, receiving sensory input from poking a cat with a finger—or implicated in pathological processes, such as epileptic seizures or sleep disorders. Brain activity is usually recorded with electro- or magnetoencephalography, abbreviated EEG and MEG, respectively. The first technique involves placing an array of electrodes on the scalp surface for measuring local electrical potentials. The second one uses sensors to record the magnetic field rather than potentials, but both measures are proxies for detecting and localizing the electrical currents in the brain.
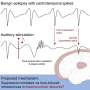
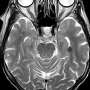
"EEG has been around for about 100 years, and some kinds of neural activity are very well-studied," the study's lead author, Senior Research Scientist Nikolay Yavich of Skoltech explained. "For example, it is fairly easy for an experienced physician to study a sleep disorder by reading raw EEG data. Other cases are more difficult. To pinpoint the precise hotspots in a patient's brain that are responsible for epileptic seizures, EEG or MEG data are combined with high-resolution MRI scans, which model the head of the patient, and processed with advanced computer algorithms. Provided that the troublesome region is accurately localized, it can then be operated without damaging the surrounding tissue to aid a patient with epilepsy when drugs do not work."
However, the MRI scans used in conjunction with brain activity maps are not always perfect. They are often corrupted by noise and other image artifacts. This leads to inaccuracies in image segmentation. According to the Skoltech researchers involved in the study, their technique is far less sensitive to such data imperfections.
"We found that when modeling neural activity on low-resolution head models, our method was up to five times more accurate than the conventional approach. While it also demands a higher computational load, the benefits seem to justify its application," Yavich commented.
This means that the method can help cognitive scientists, neurologists, and brain surgeons working with less than perfect data to understand the neurological basis underlying diseases such as epilepsy, attention deficit disorder, and autism, as well as healthy cognition processes involved in memory, sensual perception, locomotion, and more.
The technique used by the researchers is called the mixed-hybrid finite element method, or MHFEM. Its accuracy was compared against the conventional nodal finite element method, P1 FEM for short. The purpose of both methods in interpreting EEG and MEG data is to solve the equations constituting what's known as the forward problem. The methods differ in that the neural currents computed with MHFEM are always physical since they satisfy the charge conservation law, while P1 FEM does not possess this property.
More information: N. Yavich et al, Conservative Finite Element Modeling of EEG and MEG on Unstructured Grids, IEEE Transactions on Medical Imaging (2021). DOI: 10.1109/TMI.2021.3119851