Researchers identify novel pathways responsible for liver cancer
Hepatocellular carcinoma (HCC) is the most common type of primary liver cancer. It is one of leading causes of cancer-related deaths globally, with more than 700,000 new cases and 600,000 estimated HCC deaths each year. HCC occurs most often in people with chronic liver diseases such as hepatitis B, which is one of the main causes of HCC (particularly in Asia). While surgery, liver transplantation, or radiological intervention may be a viable option for early-stage disease, prognosis for advanced stage HCC remains bleak, with most patients eventually dying within 20 months after diagnosis.
A team of researchers at the Cancer Science Institute of Singapore (CSI Singapore), led by Professor Daniel Tenen and Assistant Professor Yvonne Tay, embarked on a novel study which aims to address a yet unmet clinical need. The team identified new pathways that are responsible for HCC.
Their findings were published in the journal Science Advances on 1 October 2021.
What are pseudogenes?
The CSI Singapore team focused on 'pseudogenes'—a section of chromosome that is an imperfect copy of a functional gene. A good metaphor of understanding the distinction between DNA, chromosomes, and genes is to think of the genome as a library. Our entire genome consists of many shelves of books, all of which instruct our body to produce different proteins, enzymes and other fundamental cellular materials. A gene is a single book that contains instructions for a specific product—like a single protein—while DNA sequences are the sentences in its pages. A chromosome is a shelf of books since it contains many genes, and all the shelves together combine to create the entire library (i.e. the genome). Humans usually have 23 pairs of chromosomes in each cell.
Every time a cell is created, the entire library of genomic information must be copied over to the new cell. During this process, errors in copying can occur, such as mutations. Pseudogenes describe this class of genes: genomic sequences that are similar to other genes but are defective. Continuing with the book metaphor, these are books that contain misprints or typos. Pseudogenes do not code for any protein, but they resemble genes that do, and hence are referred to as non-coding RNA (ncRNA).
Pseudogenes were once considered non-functional evolutionary relics due to their lack of coding potential, but recently, these ncRNAs have recently been linked to patient prognoses and cancer subtypes. Despite the potential clinical importance of pseudogenes, only a handful of more than 12,000 pseudogenes in humans have been characterized in cancers to date. In this study, Asst Prof Tay and her team established a previously unrecognized role for pseudogenes.
The role of pseudogenes in cancers
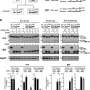
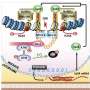
The group focused on a known gene that causes cancer, otherwise known as an 'oncogene.' The oncogene SALL4 is known to cause HCC and it contains eight pseudogenes. Since many pseudogenes are actively copied or 'transcribed' into new cells, they postulated that pseudogenes could be involved in DNA methylation—a process where a chemical methyl group (CH3) is added to the DNA strand itself. This can affect how genes are expressed—sometimes DNA methylation can repress gene expression, which is exactly what the CSI Singapore team had found.
Asst Prof Tay explained, "We found that as methylation of SALL4 increased, its expression decreased, suggesting the therapeutic potential of using DNA methylation as a regulatory mechanism to suppress the expression of SALL4 in HCC. With this interesting discovery, we decided to take a step further to investigate the correlation between methylation of a specific region in SALL4 and SALL4 expression."
The team used CRISPR technology, which allowed them to target and block gene-specific DNA methylation. They found that some SALL4 pseudogenes cause hypomethylation (the absence of CH3 methyl groups) in the CpG region. This hypomethylation—in other words, reduction of the methylation profile in the region—leads to an increased expression of the SALL4 gene with enhanced associated cellular growth. "Hence, blocking the pathways leading to the hypomethylation of the SALL4 locus could have valuable therapeutic effects on HCC patients with elevated SALL4 levels," said first author Dr. Kwon Junsu, who is a Research Fellow at CSI Singapore.
Significance and next steps
These new insights into SALL4 re-expression in HCC potentially open up avenues for the development of novel therapeutic approaches and may alter the treatment paradigm for patients.
"Moving forward, we plan to monitor the activity of pseudogenes that increase the expression of genes known to cause cancer through demethylation (i.e. the reduction of methyl groups). Through this, we hope to predict the activation of these oncogenes as well as cancer progression," said Prof Tenen.
More information: Junsu Kwon et al, Pseudogene-mediated DNA demethylation leads to oncogene activation, Science Advances (2021). DOI: 10.1126/sciadv.abg1695