Distinct gut microbial profile may identify pancreatic cancer, irrespective of stage
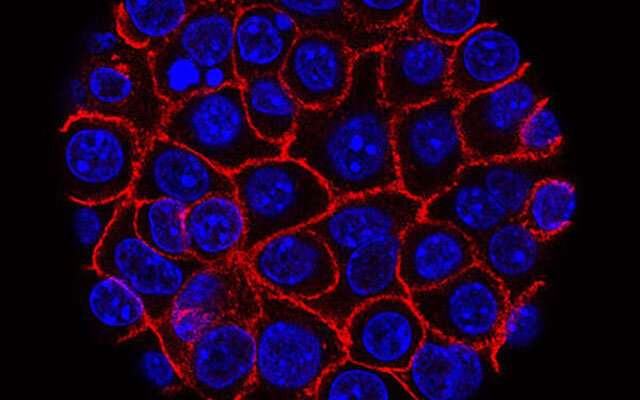
A specific panel of gut microbes may identify pancreatic cancer, irrespective of how far the disease has progressed, suggests research published online in the journal Gut.
The findings offer hope of a new non-invasive method of diagnosing this hard-to-treat disease, which currently relies on invasive procedures to detect it, say the researchers.
Pancreatic cancer is the 12th most common cancer worldwide, but is set to become even more common over the next couple of decades. The most common form of the disease is pancreatic ductal adenocarcinoma for which the outlook is poor, with fewer than 1 in 20 of those affected surviving 5 or more years.
This is largely because diagnosis is often made when the disease is advanced and because of the absence of effective treatment options. Scans, tissue specimens, and urine and blood samples are used to diagnose it. But less invasive ways of picking it up, and at an early stage, are urgently needed, say the researchers.
Recent evidence suggests that changes in the microbiome—the trillions of bacteria, fungi, and other microbes that inhabit the digestive tract—may have a role in both its development and progression.
To explore this further, the researchers analyzed 100 spit and 212 stool samples and pancreatic tissue from 57 Spanish adults newly diagnosed with the ductal form and before treatment—25 with early stage disease and 32 with advanced disease; 50 healthy people matched for age and sex; and 29 people with chronic pancreatitis, a known risk factor for pancreatic cancer.
Gut microbes were more informative than mouth microbes. After accounting for known risk factors such as smoking, drinking, obesity and diabetes, a distinct microbial profile was observed in the stool samples of people with ductal pancreatic cancer compared with people with chronic pancreatitis and those without either disease.
Machine learning techniques identified a characteristic enrichment of certain species and a relative scarcity of others. Methanobrevibacter smithii, Fusobacterium nucleatum, Alloscardovia omnicolens, Veillonella atypica and Bacteroides finegoldii were abundant in the stool samples of the cancer patients while Faecalibacterium prausnitzii, Bacteroides coprocola, Bifidobacterium bifidum or Romboutsia timonensis were depleted.
This microbial profile consistently identified patients with the disease, irrespective of how far it had progressed suggesting that characteristic microbiome signatures emerge early on and that the stool microbiome might pick up early stage disease, say the researchers.
Predictive ability was assessed using the area under the receiver operating characteristic (AUROC) curve, a graphic representation of how well a test identifies and excludes a given condition. An area beneath the ROC curve of 0.5 corresponds to random chance;1 equals perfect accuracy. In this case, the AUROC curve reached 0.84.
Accuracy increased still further to 0.94 when the microbial profile was combined with blood levels of carbohydrate antigen 19-9, a substance indicative of pancreatic cancer, but also various other conditions, and the only current non-invasive test, approved by the US drugs regulator, the Food and Drug Administration (FDA), for monitoring pancreatic cancer progression.
The predictive ability of the microbial profile was validated in a separate group of 76 German people, 44 of whom had pancreatic ductal cancer and 32 of whom didn't.
It was then validated against publicly available data from 25 studies involving 5792 samples covering 9 different health conditions, including other cancers and type-2 diabetes, a risk factor for pancreatic cancer.
The microbial profile of mouth, stool, and pancreatic tissue samples of patients with ductal pancreatic cancer were similar, suggesting that they may be closely linked.
"Our data are strictly observational and cross-sectional," highlight the researchers. "Nevertheless, there are strong indications that the identified fecal microbiome shifts are not merely a consequence of impaired pancreatic function or systemic effects thereof, although indirect effects cannot be ruled out," they add.
And they go as far to say that given previous research on a possible link between pancreatic cancer and the gut microbiome, "we believe that the presented panel of [ductal pancreatic cancer]] associated bacterial species may be relevant beyond their use for diagnosis, providing promising future entry points for disease prevention and therapeutic intervention."
In a linked commentary, Drs. Rachel Newsome and Christian Jobin of the University of Florida, caution: "Although promising, these findings have limited clinical value due to the cross-sectional nature of the study, and thus the predictive markers will need to be tested using a prospective cohort before reaching a conclusion on their clinical impact."
Further research would be needed to find out if the microbial profile is specific for pancreatic cancer and not shared with other types of cancer, they add.
Notwithstanding, the research "represents an important contribution toward the generation of predictive markers for [ductal pancreatic cancer] and highlights the key role of microbiota in cancer surveillance……"and represents significant progress for non-invasive cancer detection," they write.
More information: A faecal microbiota signature with high specificity for pancreatic cancer, Gut (2022). gut.bmj.com/lookup/doi/10.1136/gutjnl-2021-324755