Researchers develop new computational tool to interpret clinical significance of cancer mutations
Researchers at Children's Hospital of Philadelphia (CHOP) have developed a new tool to help researchers interpret the clinical significance of somatic mutations in cancer. The tool, known as CancerVar, incorporates machine learning frameworks to go beyond merely identifying somatic cancer mutations and interpret the potential significance of those mutations in terms of cancer diagnosis, prognosis, and targetability. A paper describing CancerVar was published today in Science Advances.
"CancerVar will not replace human interpretation in a clinical setting, but it will significantly reduce the manual work of human reviewers in classifying variants identified through sequencing and drafting clinical reports in the practice of precision oncology," said Kai Wang, Ph.D., Professor of Pathology and Laboratory Medicine at CHOP and senior author of the paper. "CancerVar documents and harmonizes various types of clinical evidence including drug information, publications, and pathways for somatic mutations in detail. By providing standardized, reproducible, and precise output for interpreting somatic variants, CancerVar can help researchers and clinicians prioritize mutations of concern."
"Somatic variant classification and interpretation are the most time-consuming steps of tumor genomic profiling," said Marilyn M. Li, MD, Professor of Pathology and Laboratory Medicine, Director of Cancer Genomic Diagnostics and co-author of the paper. "CancerVar provides a powerful tool that automates these two critical steps. Clinical implementation of this tool will significantly improve test turnaround time and performance consistency, making the tests more impactful and affordable to all pediatric cancer patients."
The growth of next-generation sequencing (NGS) and precision medicine has led to the identification of millions of somatic cancer variants. To better understand whether those mutations are related to or impact the clinical course of disease, researchers have established several databases that catalogue these variants. However, those databases did not provide standardized interpretations of somatic variants, so in 2017, the Association for Molecular Pathology (AMP), American Society of Clinical Oncology (ASCO), and College of American Pathologists (CAP) jointly proposed standards and guidelines for interpreting, reporting, and scoring somatic variants.
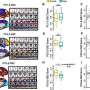
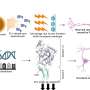
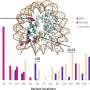
Yet even with these guidelines, the AMP/ASCO/CAP classification scheme did not specify how to implement these standards, so different knowledge bases were providing different results. To solve this problem, the CHOP researchers, including CHOP data scientist and co-senior author of the paper Yunyun Zhou, Ph.D., developed CancerVar, an improved somatic variant interpretation tool using command-line software called Python with an accompanying web server. With a user-friendly web server, CancerVar includes clinical evidence for 13 million somatic cancer variants from 1,911 cancer census genes that were mined through existing studies and databases.
In addition to including millions of somatic mutations, whether of known significance or not, the tool uses deep learning to improve clinical interpretation of those mutations. Users can query clinical interpretations for variants using information such as the chromosome position or protein change and interactively fine-tune how specific scoring features are weighted, based on prior knowledge or additional user-specified criteria. The CancerVar web server generates automated descriptive interpretations, such as whether the mutation is relevant for diagnosis or prognosis or to an ongoing clinical trial.
"This tool shows how we can use computational tools to automate human generated guidelines, and also how machine learning can guide decision making," Wang said. "Future research should explore applying this framework to other areas of pathology as well."
More information: Quan Li et al, CancerVar: An artificial intelligence–empowered platform for clinical interpretation of somatic mutations in cancer, Science Advances (2022). DOI: 10.1126/sciadv.abj1624. www.science.org/doi/10.1126/sciadv.abj1624