Risk for miscarriage and failure of in vitro fertilization can be predicted with genomic data
A woman's risk of suffering one of the most common types of miscarriages can be predicted based on a specialized analysis of her genome, according to a Rutgers University study.
Scientists said this insight could allow patients and clinicians to make better-informed decisions regarding reproductive choices and fertility treatment plans.
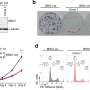
Reporting in the science journal Human Genetics, Rutgers researchers describe a technique combining genomic sequencing with machine-learning methods to predict the possibility a woman will undergo a miscarriage because of egg aneuploidy—a term describing a human egg with an abnormal number of chromosomes.
Infertility is a major reproductive health issue that affects about 12 percent of women of reproductive age in the U.S. Aneuploidy in human eggs accounts for a significant proportion of infertility, causing early miscarriage and in vitro fertilization (IVF) failure.
Recent studies have shown that genes predispose certain women to aneuploidy, but the exact genetic causes of aneuploid egg production have remained unclear. The Rutgers study is the first to evaluate how well individual genetic variants in the mother's genome can predict a woman's risk of infertility.
"The goal of our project was to understand the genetic cause of female infertility and develop a method to improve clinical prognosis of patients' aneuploidy risk," said Jinchuan Xing, an author of the study and an associate professor in the genetics department at the Rutgers School of Arts and Sciences. "Based on our work, we showed that the risk of embryonic aneuploidy in female IVF patients can be predicted with high accuracy with the patients' genomic data. We also have identified several potential aneuploidy risk genes."
Working with Reproduction Medicine Associates of New Jersey, an IVF clinic in Basking Ridge, N.J., the scientists were able to examine genetic samples of patients using a technique called "whole exome sequencing," which allows researchers to home in on the protein coding sections of the vast human genome. Then they created software using machine learning, an aspect of artificial intelligence in which programs can learn and make predictions without following specific instructions. To do so, the researchers developed algorithms and statistical models that analyzed and drew inferences from patterns in the genetic data.
As a result, the scientists were able to create a specific risk score based on a woman's genome. The scientists also identified three genes—MCM5, FGGY and DDX60L—that when mutated, are highly associated with a risk of producing eggs with aneuploidy.
While age is a predictive factor for aneuploidy, it is not a highly accurate gauge because aneuploidy rates within individuals of the same age can vary dramatically. Identifying genetic variations with more predictive power arms women and their treating clinicians with better information, Xing said.
"I like to think of the coming era of genetic medicine when a woman can enter a doctor's office, or in this case, perhaps, a fertility clinic with her genomic information, and have a better sense of how to approach treatment," Xing said. "Our work will enable such a future."
More information: Siqi Sun et al, Predicting embryonic aneuploidy rate in IVF patients using whole-exome sequencing, Human Genetics (2022). DOI: 10.1007/s00439-022-02450-z