New research maps possible molecular origins of kidney disease
After mapping the genetic underpinning of kidney function in 1.5 million people and about 60,000 kidney cells that are the microscopic mechanisms of gene regulation, researchers at the Perelman School of Medicine at the University of Pennsylvania found that more than 500 genes likely contribute to kidney disease development. Multiple genes that play a key role in kidney detoxification, including SLC47A1, have been identified. The good news is that close to 100 of the 500 genes might be able to be targeted by various pharmaceuticals already approved by the Food and Drug Administration (FDA). The findings point to the underlying genetics of kidney function and can lead to future research into possible therapeutic targets to treat kidney disease and its development. The research was published in Nature Genetics.
"Single cell level analytical tools now allow us to dive deeper into the mechanisms at play by uncovering genetic variation in all cells in the body, and then we can look for links to genetics of diseases and illnesses," said Katalin Susztak, MD, Ph.D., lead investigator and a professor of Nephrology and Genetics at Penn. "Highlighting the links and exploring cause and effect will offer greater understanding of the human kidney and uncover potential ways to treat those who struggle with kidney issues."
An estimated 37 million people in the United States have kidney disease, and mortality from kidney disease has risen by more than 40% in the last two decades, making it one of the fastest-growing causes of death. Roughly a million people die of kidney failure worldwide each year. Despite the major personal and economic burden, few new therapeutics have been registered to treat or cure kidney disease over the last 40 years.
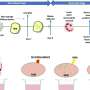
Using genetic information from more than 1.5 million participants around the world and a basic-science technique of mapping cells known as single-cell sequencing, scientists at Penn implicated important cell types for different disease conditions such as the proximal tubules for kidney disease and collecting duct principal cells for high blood pressure.
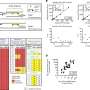
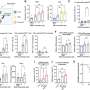
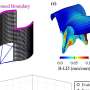
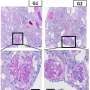
Next, the researchers characterized gene expression and gene regulation in hundreds of human kidney samples and analyzed changes in more than 60,000 human kidney cells. The team used sophisticated computational and statistical methods to generate the most comprehensive maps to uncover the genes, cell types and mechanism of kidney dysfunction.
"While there may be many origins of kidney dysfunction in human kidneys, our studies specifically highlight the role of SLC471A gene," said Hongbo Liu, Ph.D., a postdoctoral fellow in Susztak' s lab. "This gene carries different toxins. Changes in SLC47 might make the kidneys of people more or less sensitive to toxin mediated injury and kidney disease."
The study authors want to continue to look into specific genes to understand their role in kidney diseases and say that their study serves as a springboard for researchers to test different pharmaceuticals against these genetic variants and their deleterious effects.
"There may come a time many years from now when patients who have these genetic variants can receive treatment before kidney disorders arise," Susztak said.
More information: Hongbo Liu et al, Epigenomic and transcriptomic analyses define core cell types, genes and targetable mechanisms for kidney disease, Nature Genetics (2022). DOI: 10.1038/s41588-022-01097-w
Datasets produced in this study are available at susztaklab.com