Protein signature identifies those at highest risk from severe infection
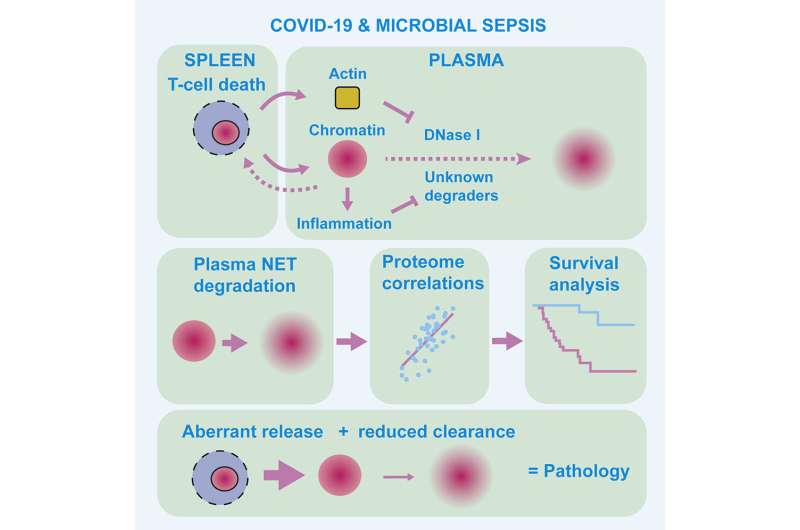
Scientists at the Francis Crick Institute have uncovered how a build-up of DNA molecules can impact survival chances in people with severe COVID-19 or sepsis, and have developed a tool that could be used to identify the patients at most risk.
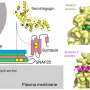
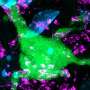
Chromatin is a complex molecule of DNA and proteins, that forms chromosomes within the nucleus of cells and is vital for regulating gene expression. When cells die, chromatin is released into the surroundings and can cause inflammation and cytotoxicity. A process known as chromatin clearance is needed to remove extracellular chromatin and protect against severe disease.
In their research, published in Immunity today, the team aimed to understand how infections affect chromatin clearance, by studying the protein levels and enzyme activity in cases of severe disease.
Working with clinicians in Germany and Switzerland, and protein analysis experts led by Professor Markus Ralser, the team studied samples taken from 63 people hospitalized with severe COVID-19 pneumonia. All individuals fit the most severe category of disease, as defined by the World Health Organization.
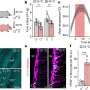
They found that chromatin clearance was hindered in all cases and, and when this process was impacted the most, these people were less likely to survive.
Further analysis into the cause of this chromatin build-up highlighted that a group of enzymes, called DNAses, were less active. These enzymes help to break down DNA molecules, including chromatin. The activity of the enzymes was impacted by another molecule, called actin, which is released when cells die.
The researchers also studied blood plasma samples taken from 35 people with microbial sepsis. A build-up of chromatin was also seen in this group, and this was highest in people with high levels of actin in the blood.
Using this information, the team were able to develop a 'proteomic profile', a signature of protein levels and enzyme activity in the blood, that characterized the most severe and high-risk cases of infection. With further development, this signature could be used by doctors to help distinguish patients who might require additional treatment.
Iker Valle Aramburu, first author and postdoctoral training fellow at the Crick, says, "We know that an accumulation of chromatin can be harmful, but the underpinning mechanisms have been poorly understood. We've helped answer this mystery here and also shown that the most severe cases of infection are characterized by both a release of chromatin alongside defective chromatin clearance."
Veni Papayannopoulos, senior author and principal group leader at the Crick, says, "Our results might also help identify healthy people who are at higher risk of severe disease before they become infected. We saw that chromatin degrading enzymes are also less active in cases of healthy people who have low levels of inflammation and a high risk for cardiovascular disease. This could put this group at higher risk of massive build-up of chromatin upon infection, as the process of chromatin clearance is already weakened."
The researchers will continue this work, with an aim of developing algorithms capable of scanning large data sets to identify key activity-associated patterns in protein levels.
More information: Iker Valle Aramburu et al, Functional proteomic profiling links deficient DNA clearance with increased mortality in individuals with severe COVID-19 pneumonia, Immunity (2022). DOI: 10.1016/j.immuni.2022.11.007