This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New SARS-CoV-2 target could boost immunity against all coronaviruses
Researchers searching for new therapeutic targets in the SARS-CoV-2 virus have identified a potential Achilles' heel that exists in all coronaviruses, according to a study published today in eLife.
The research could aid the development of more powerful antibody drugs and vaccines against the virus that currently causes COVID-19 and might also protect against emerging coronaviruses that could cause future pandemics.
Most vaccines and antibody-based treatments for COVID-19 neutralize the SARS-CoV-2 virus by disrupting interactions between the protein spike on the virus and the ACE2 receptor on human cells, which the virus hijacks to gain entry. But mutations in the spike protein mean that emerging variants of SARS-CoV-2 can escape the human antibody response, so that treatments do not work and vaccinated individuals experience breakthrough infections. One way to address this issue is to target treatments and vaccines against parts of the spike protein that the virus needs for survival and cannot mutate.
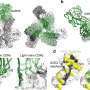
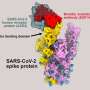
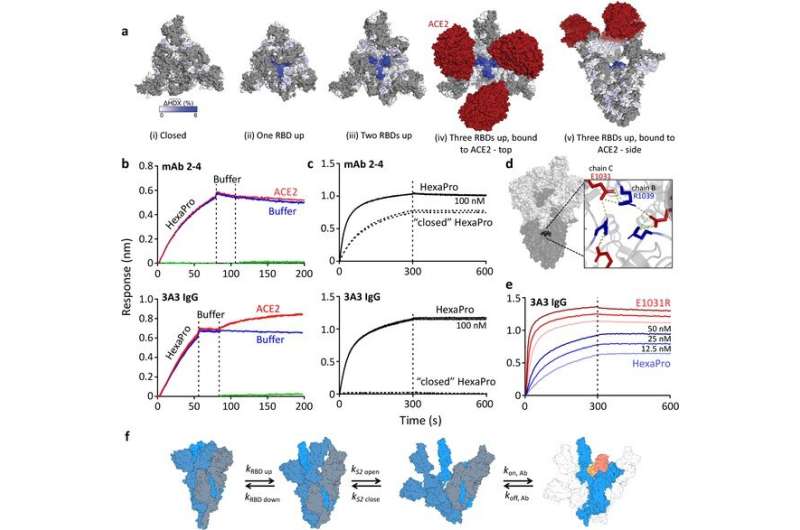
The SARS-CoV-2 spike protein is made up of two subunits—called S1 and S2. The S1 subunit contains a region that binds to the ACE2 receptor, while the S2 subunit allows the virus to fuse with the membrane of the cell it is gaining access to. Most mutations in the spike protein affect its S1 domain, but the S2 domain is highly conserved across all seven human coronaviruses, suggesting that it could be a good target for therapeutic antibodies and vaccines.
"We know that S2-directed antibodies are produced following a COVID-19 infection, and similar subunits in flu and HIV are also targeted by the body's antibodies," explains Rui Silva, a former graduate student at The University of Texas at Austin, U.S., who led the work in close collaboration with fellow graduate student Yimin Huang and Senior Scientist Annalee Nguyen.
"However, we know very little about the antibodies that bind the S2 subunit. Fewer than five percent of the roughly 7,000 known anti-SARS-CoV-2 antibodies bind S2 and only two classes of S2 binding antibodies have been characterized in detail. We have identified a third class of S2 antibody that binds a highly conserved part of the S2 subunit."
The team isolated the antibody by immunizing mice with the S2 subunit from the related Middle Eastern respiratory syndrome virus and collected the antibodies produced that bound spike from this virus as well as SARS-CoV-2. They then analyzed the most promising anti-S2 antibody further.
They found that this antibody binds a hinge in the S2 subunit that plays a crucial role when the subunit changes its shape during the virus's fusion with the human cell membrane. Further analysis showed that access to the point where the antibody binds to this hinge (called the epitope) depended on the shape-shifting dynamics of the overall spike protein; when the spike is bound to the ACE2 receptor, the epitope on the hinge is accessible.
The team then conducted a series of experiments to see if targeting this hinge epitope had any impact on viral activity. They found that although S2-directed antibodies could prevent the spike protein's ability to fuse viral and human cell membranes, they were less potent at neutralizing the virus altogether.
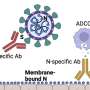
However, as well as directly neutralizing viruses, antibodies also indirectly kill virus-infected cells, by triggering other processes. In one experiment, S2-directed antibodies could trigger human natural killer cells to destroy SARS-CoV-2 infected cells. In another, the S2 antibodies caused cells called monocytes to gobble up and destroy the infected cells. This suggests that although S2 antibodies might not be strong enough to directly block infection, they could be used to help boost or protect an immune response.
"We have identified an epitope in the S2 subunit of SARS-CoV-2 that is highly conserved across all pathogenic coronavirus strains," concludes senior author Jennifer Maynard, ZD Bonner Professor of Chemical Engineering at The University of Texas at Austin. "Although targeting this epitope alone is unlikely to be potent enough as a treatment or vaccine, therapeutic strategies that can enhance access to this epitope could allow existing human antibodies to more effectively promote viral clearance through other antibody-directed cell-killing processes."
More information: Rui P Silva et al, Identification of a conserved S2 epitope present on spike proteins from all highly pathogenic coronaviruses, eLife (2023). DOI: 10.7554/eLife.83710