This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Team successfully generates mouse models for two subtypes of multiple myeloma
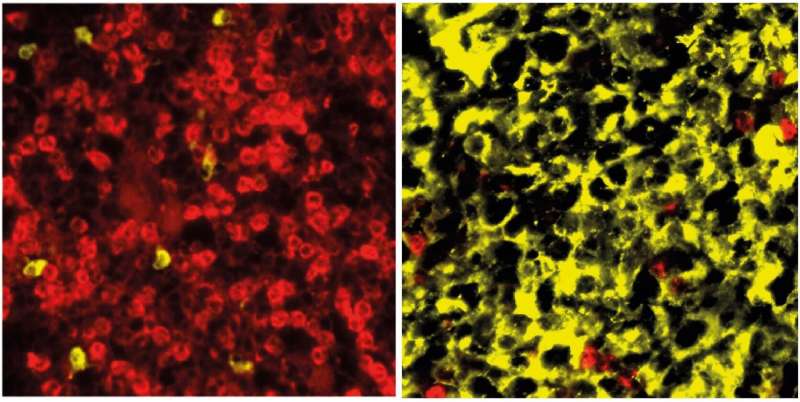
B lymphocytes—also known simply as B cells—play a central role in the immune system. If pathogens enter the body, B cells are activated and develop into plasma cells, which then release antibodies. One important step in this process is the germinal center reaction. If the B cells' maturation into plasma cells is disrupted, multiple myeloma can develop—one of the most common blood cancers. This disease has a variety of subtypes and is not yet curable.
Multiple myelomas develop very slowly and in several stages. The process is initiated by spontaneous genetic aberrations that occur during the germinal center reaction and influence the process of B-cell maturation. The preliminary stage of the disease is called monoclonal gammopathy of undetermined significance (MGUS)—a benign precursor that causes no symptoms.
The only biomarker is an increased concentration of the antibody secreted by the plasma cells in the blood. It takes further genetic changes in the plasma cells for the line between MGUS and malignant cancer to be irreversibly crossed.
Models exactly mimic the human disease
Previous mouse models have not been able to accurately represent the different genetic subtypes of myeloma. A team led jointly by renowned B-cell researcher Professor Klaus Rajewsky and lymphoma expert Dr. Martin Janz has now succeeded in doing just that. In the journal PNAS, they present novel mouse models that precisely replicate two subtypes of human multiple myeloma.
"We have also been able to show that the interaction of several genetic aberrations is a decisive factor in the development of the disease," says Janz, head of the Biology of Malignant Lymphomas Lab at the Experimental and Clinical Research Center (ECRC), a joint institution of the Max Delbrück Center and Charité—Universitätsmedizin Berlin.
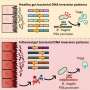
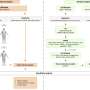
The researchers started by establishing three different groups of transgenic mice, each carrying just one genetic modification—an extra copy of the genes that encode either cyclin D1, MMSET, or Ikk2. Cyclin D1 regulates cell cycle progression, and the incorrect activation of its encoding gene due to an aberration promotes increased cell division. MMSET is a histone methyltansferase that regulates the accessibility of DNA.
Overexpression of its encoding gene profoundly changes the epigenetic pattern of the cell and enhances its susceptibility to undergo malignant transformation. Ikk2 activates a component of the NF-κB signaling pathway, which plays an important role in cell growth and immune response. The frequent activation of this signaling chain is a distinguishing characteristic of multiple myeloma.
Multiple myeloma mostly occurs in old age
In a second step, the scientists crossed the cyclin D1 and MMSET mice with Ikk2 mice and selected the offspring with the desired genetic attributes—i.e., cyclin D1 + Ikk2 and MMSET + Ikk2. They then mated these with another mouse strain, which allowed the modified genetic information to be activated only in the B cells and only as part of the germinal center reaction.
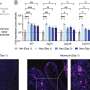
"It was striking to see how the primary modification in the mouse model—through overexpression of either cyclin D1 or MMSET—really shapes the profile of the disease subtype," says Rajewsky.
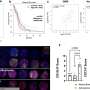
It took 70 to 90 weeks for the experimental mice to develop full-blown multiple myeloma—a long time in the life of a mouse. Though this timeline complicates the experiments, it does accurately mimic the development of the disease in humans: for us, too, multiple myeloma tends to emerge later in life and often takes years to progress to the malignant stage. It is estimated that up to five percent of all people over the age of seventy have the benign precursor MGUS.
"Our models make it clear that multiple myeloma only develops when several genetic aberrations occur together," Janz explains. "Mice that were only transgenic with regard to cyclin D1 or MMSET and did not also carry modified Ikk2 did not develop the disease."
An important starting point for further tests
Although the symptoms are similar—elevated calcium levels, anemia, fatigue, increased susceptibility to infection, renal insufficiency, bone damage—multiple myeloma subtypes in humans differ with regard to the nature of the genetic changes, gene expression profiles, and prognosis.
"Our models provide an important foundation for investigating the differences and similarities between the various subgroups and will help us to develop more specific, individualized therapeutic strategies in the long term," says Dr. Wiebke Winkler, the study's lead author.
The researchers now want to use the novel mouse models to identify the subgroups' "genetic Achilles heels," as Janz puts it. In addition, they want to activate the B cells in the animal models in an even more targeted way and to introduce more secondary genetic modifications in the mouse genome. "After all, Ikk2 is not the only driver of the disease," stresses Janz.
More information: Wiebke Winkler et al, Mouse models of human multiple myeloma subgroups, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2219439120