Photo: NIAID
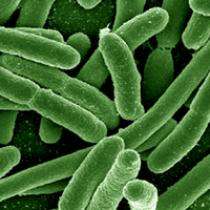
Although they're among the simplest organisms, bacteria are some of the most creative life forms on earth. Just ask molecular biologist Chobi DebRoy, director of Penn State's Gastroenteric Disease Center. "We receive bacterial samples from all over the world -- from veterinary clinics, medical centers, zoos, farms," DebRoy said. Her lab works to identify disease-causing strains of E. coli and to determine whether they are antibiotic-resistant.
Bacteria such as Escherichia coli can alter their genetic makeup with astonishing rapidity. They do this, said DebRoy, "through mutations in normal cellular genes, by acquiring foreign-resistance genes from the environment or by a combination of those mechanisms."
Consider an animal -- a human or a cow, say -- with the usual suite of bacteria in the gastroenteric system (the stomach and intestines). If a strain of bacteria starts to overcome the body's immune system, a doctor or a veterinarian may administer an antibiotic to fight the infection. But if not enough of the antibiotic is given or if treatment is stopped too soon, the drugs may fail to kill all of the bacteria. Low levels of antibiotics are sometimes given to patients, so they won't develop infections, and to animals used for milk and meat production, so they won't get sick and so they'll grow faster. These "subtherapeutic levels" of antibiotics also may help antibiotic-resistant bacteria to emerge.
Some antibiotics work by binding to the cell walls of bacteria, breaking down the walls and killing the microbes. Resistant bacteria change their cell walls slightly, so the antibiotics cannot attach or they produce enzymes to disable the antibiotics. Other drugs target the ribosomes of bacteria -- particles of RNA and protein inside the unicellular organisms that help in the synthesis of proteins necessary for reproduction. Some bacteria evolve slight changes in their ribosomes so that the drugs cannot bind to the particles.
A strain of E. coli may acquire genes from a highly resistant Salmonella strain in the same environment. The genetic material may arrive through plasmids -- circular, double-stranded units of DNA, or it may be transferred inside a phage, a virus that infects bacteria. Recently, scientists have discovered a previously unknown vehicle, dubbed a "gene cassette" or an "integron," that bacteria use to acquire and disseminate the genes conferring antibiotic resistance.
When a bacterial sample comes in the door at the Gastroenteric Disease Center, DebRoy and her colleagues use serotyping analysis to examine the kinds of O and H antigens present on the surface of the bacteria -- antigens that spell the difference between a bacterium that's benign or one that's virulent. "Bacterial genes have been widely mapped, cloned and sequenced," noted DebRoy. "Scientists have determined the sequences of the genes that encode for virulence. We can detect the presence of those genes by using a technique called Polymerase Chain Reaction."
Recently the center has participated in surveillance studies to detect the presence of pathogenic E. coli in meat, fruits and vegetables. The center helps petting zoos and other facilities draw up guidelines for human/animal interactions. "Our center is one of the largest repositories of E. coli strains in North America," said DebRoy. "We have more than 70,000 isolates, collected over the last 40 years from a variety of hosts, including humans, cows, pigs, birds and the greater environment." Scientists at Penn State and elsewhere use samples from the collection to establish relatedness between strains of bacteria -- and to chart the frightening trend toward antimicrobial drug resistance shown in recent years by these adaptable life forms.
Source: By Charles Fergus, Research Penn State