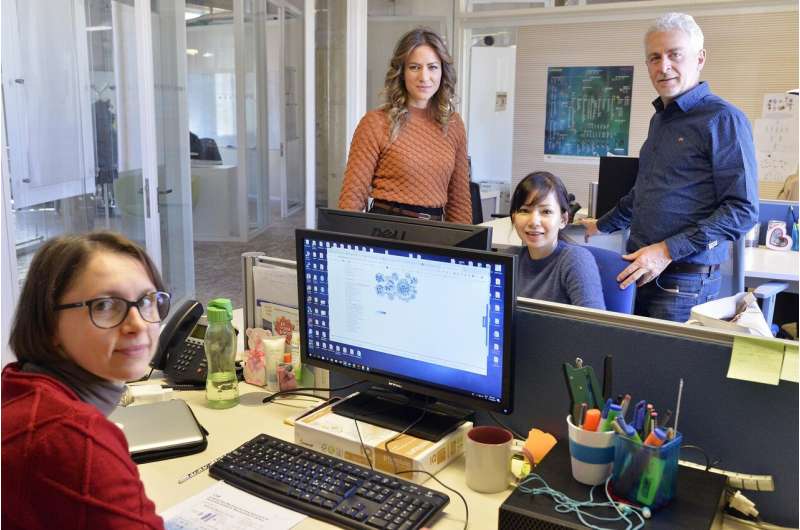
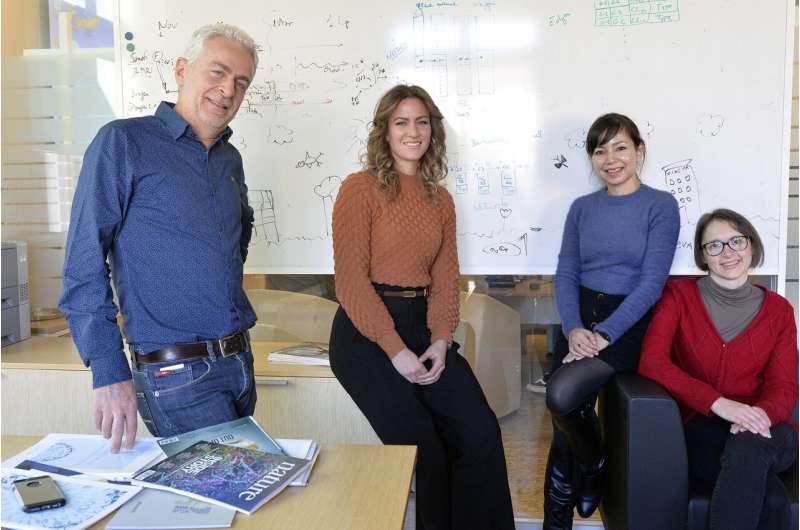
The research group at The Microsoft Research - University of Trento Centre for Computational and Systems Biology. Silvia Parolo, Pranami Bora, Lorena Leonardelli, Enrico Domenici. Credit: ©AlessioCoser
New drugs can be developed from old ones. This revolutionary pharmacological approach was confirmed by a study published today in Nature Communications that was conducted by two major research centers of the University of Trento: one at Cosbi (Fondazione The Microsoft Research—University of Trento Centre for Computational and Systems Biology), which is specialized in IT and big data, and one at Cibio Department, which is focused on biology and genomics.
With a new algorithm designed by Cosbi, the researchers first analyzed the existing drugs, particularly their molecular properties, to see whether they could be used to treat patients with other conditions that are difficult or impossible to treat. This technique, known as drug repositioning, has been used in the past, but it has become even more effective today thanks to quick and systematic analysis of vast amounts of data.
"We have achieved promising results," said Enrico Domenici, president of Cosbi. "We have tested the new algorithm to search for new therapies to treat metabolic syndrome, a condition that increases the risk of heart disease and type 2 diabetes, and which is characterized by obesity, elevated levels of cholesterol and/or triglycerides, high blood pressure and diabetes. We identified the mutated genes that are responsible for these alterations and then searched pharmaceutical databases for already approved molecules that could interact with the networks that these genes create in the adipose tissue, liver and muscles. To carry out such a complex study, we adopted a new computational approach that measures the 'distance' between the proteins targeted by the drug and those found in the networks affected by the disease. That is how we found a new purpose for an already existing drug: Ibrutinib, which was originally developed to treat completely different conditions, and specifically B-cell tumors like mantle cell lymphoma, chronic lymphocytic leukemia, and Waldenström macroglobulinemia. Other studies on this drug are under way to determine if it could be used to treat autoimmune diseases."
The results of the computational analysis were validated at Cibio on zebrafish larvae to confirm the response to the drug. The head of the research team at Cibio department, Maria Caterina Mione, explained how it worked: "When we tested this drug in the laboratory we saw that the devastating impact of obesity caused by a high-fat diet could be limited. The drug succeeded in limiting the inflammatory condition associated with lipid accumulation. Testing the effectiveness of a drug that is already on the market makes it possible to skip a number of long and complex stages of the approval process for new medical products, because their tolerance and safety have already been demonstrated. Our data are just a start and more in-depth studies and clinical tests are required, but they demonstrate that combining experience in big data analysis and the ability to develop models for biological validation can boost drug repurposing research."
The research group at The Microsoft Research - University of Trento Centre for Computational and Systems Biology. Silvia Parolo, Pranami Bora, Lorena Leonardelli, Enrico Domenici. Credit: ©AlessioCoser
Finding out that a market-approved drug can potentially be used to treat metabolic syndrome is great news. The incidence of the disease and seriousness of its consequences on health (cardiovascular diseases and diabetes complications) have made metabolic syndrome a societal problem in recent decades, especially in wealthy societies, where many people have a diet high in sugars and fats and a sedentary lifestyle. In particular, the disease affects people over 50 years of age, about 30 percent of the male population, and 35 to 40 percent of women. Metabolic syndrome becomes more common in women after menopause.
The study skipped a few phases of the drug development process and proved effective for repurposing a drug that was originally developed to treat other conditions. However, more needs to be done in the years to come as the research work will move to clinical trials.
More information: Karla Misselbeck et al. A network-based approach to identify deregulated pathways and drug effects in metabolic syndrome, Nature Communications (2019). DOI: 10.1038/s41467-019-13208-z
Journal information: Nature Communications
Provided by Università di Trento