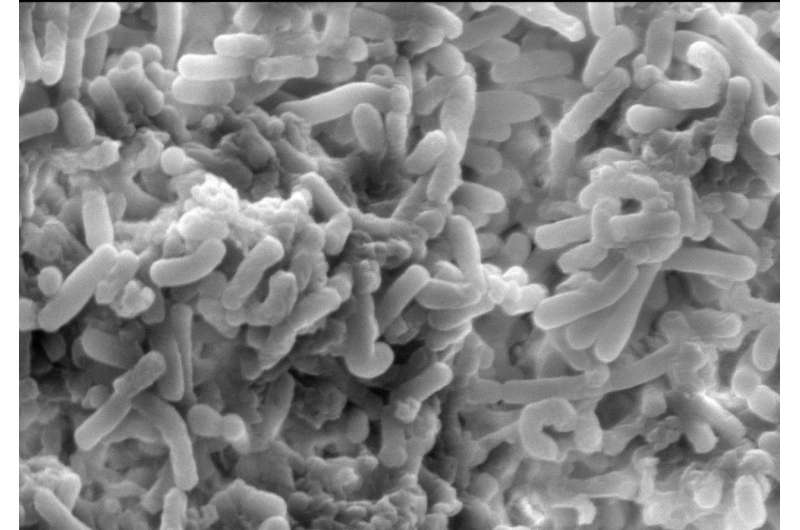
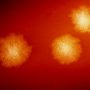
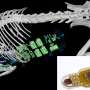
Scanning electron microscopy image of Clostridium difficile community. Credit: Kerrie Venner (UCL Institute of Neurology) and Lisa Dawson (LSHTM)
The bacterium Clostridium difficile, which is responsible for the majority of antibiotic-associated diarrhea outbreaks worldwide, produces a unique compound called p-cresol to gain a competitive advantage over natural protective gut bacteria. The findings were reported by Lisa Dawson and Team at the London School of Hygiene and Tropical Medicine and colleagues on September 11 in the open-access journal PLOS Pathogens.
Antibiotics disrupt the natural protective gut flora, rendering people susceptible to C. difficile infection, which leads to potentially life-threatening disease and complications. Currently, there is a great need to better understand how C. difficile is able to influence the gut microbiota and disrupt intestinal homeostasis. One clue to this question is that C. difficile is one of only 18 intestinal commensal species known to produce p-cresol, which inhibits the growth of a broad range of microorganisms but only affects C. difficile at relatively high concentrations.
Dawson and colleagues now show for the first time that the production of p-cresol by C. difficile confers a fitness advantage over other gut bacteria. Experiments in mice showed that p-cresol selectively targets certain bacteria in the gut and disrupts their ability to grow, leading to significant alterations in the intestinal microbiota. Moreover, mutant C. difficile strains that could not produce p-cresol were less capable of recolonizing the gut after initial infection. The findings suggest that p-cresol production contributes to C. difficile survival and pathogenesis.
Dawson notes, "we have identified that the major gut pathogen Clostridium difficile produces the bacteriostatic agent para-cresol which helps control the intestinal microbiota and provides C. difficile with a competitive growth advantage particularly after the consumption of antibiotics. This unique attribute of the pathogen may provide a novel drug target to reduce C. difficile infection."
More information: PLOS Pathogens (2018). journals.plos.org/plospathogen … journal.ppat.1007191
Journal information: PLoS Pathogens
Provided by Public Library of Science