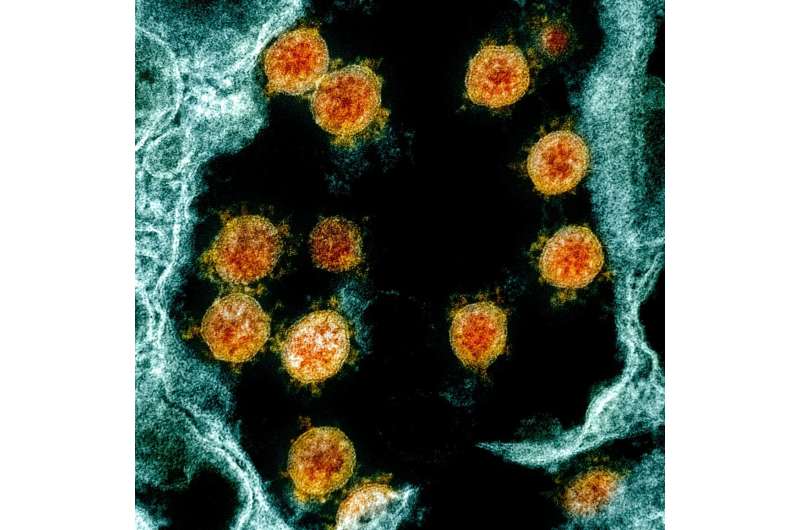
Colorized transmission electron micrograph shows SARS-CoV-2 virus particles (orange), isolated from a patient. Credit: NIAID via NIH.
The symptoms and side effects of COVID-19 are scattered across a diagnostic spectrum. Some patients are asymptomatic or experience a mild immune response, while others report significant long-term illnesses, lasting complications, or suffer fatal outcomes.
Three researchers from the Georgia Institute of Technology and one from Emory University are trying to help clinicians sort through these factors and spectrum of patient outcomes by equipping healthcare professionals with a new "decision prioritization tool."
The team's new artificial intelligence-based tool helps clinicians understand and better predict which adverse effects their COVID-19 patients could experience, based on comorbidities and current side effects—and, in turn, also helps suggest specific Food and Drug Administration-approved (FDA) drugs that could help treat the disease and improve patient health outcomes. The researcher's latest findings are the focus of a new study published October 21 in Scientific Reports.
Artificial intelligence, protein drivers, and '24 out of 26 clinical manifestations' of COVID-19
The team's new methodology, or tool, is called MOATAI-VIR (Mode Of Action proteins & Targeted therapeutic discovery driven by Artificial Intelligence for VIRuses.) Researchers report it predicts 24 out of 26 major clinical manifestations of COVID-19 and their underlying disease-protein-pathway relationships.
Those clinical manifestations cover a range of issues including acute respiratory distress, blood clotting issues, cytokine storms, low blood oxygen and white blood cell counts, and even bone marrow failure. The commonly reported loss of smell or taste, along with unusual neurological symptoms such as "brain fog" have received widespread attention—as have considerations for patients who have previous health problems, or comorbidities, that could place them in higher risk categories related to COVID-19.
"It's still the question of, what's causing the side effects?" says Jeffrey Skolnick, professor and Mary and Maisie Gibson Chair in the School of Biological Sciences, and corresponding author for the study. "So, you lost sense of smell and got brain fog—and another (patient) had respiratory distress, and another can't remember the day of the week. What we've identified are the possible mode of action drivers for these various conditions, which is now setting the stage for who's getting what side effects."
Skolnick, also Georgia Research Alliance Eminent Scholar in Computational Systems Biology, notes that it makes sense to predict the side effects based on protein interactions.
"Humans are molecular machines, and presumably there are biological and physical rules to dictate our responses," said Skolnick. "We basically built an AI-based approach which was designed given the interactive set of proteins in humans which interact with the [novel] coronavirus," he adds. "We then asked ourselves, 'Could we predict, based on biochemical pathways, which interactive proteins are associated with side effects?'"
Joining Skolnick from the School of Biological Sciences are Ph.D. student Courtney Astore and senior research scientist Hongyi Zhou, both from the Center for the Study of Systems Biology. Joshy Jacob of the Department of Microbiology and Immunology in the Emory Vaccine Center at the Emory School of Medicine also worked on the study.
MOATAI-VIR methodology
Skolnick explains that most known diseases are due to the "malfunction and interaction of many proteins," and notes that it's a collective effect—a "many-targeted protein effect." His team's new AI methodology is identifying as many targets as possible of an interacting nature to predict a complex response from a complex set of interactions.
It's also well-understood in the medical community that comorbidities—existing and chronic health factors such as diabetes, obesity, autoimmune disorders, and other conditions that affect the immune system—can play an outsized a role in risk factors related to COVID-19. Skolnick says those comorbidities can be plugged into the team's algorithm, too.
"Alzheimer's and hyperthyroidism are strongly correlated—as is diabetes. There are six to eight (COVID-19) comorbidities with a patient that has Alzheimer's," Skolnick explains. "It's not just old age—it's much more complicated."
The MOATAI-VIR methodology helps identify the common proteins of the comorbidities in relation to the parent disease. A clinician can then target the diseases with drugs. Researchers report that this specific methodology had 72% success in 123,146 drug-indication pair predictions found by Skolnick's team.
"For a given disease, we prioritize them by the proteins that are most in common with the comorbid diseases to the given disease, giving rise to the particular complication, such as respiratory failure. This identifies the putative (assumed) driver proteins for the given complication," he says. "Then we select repurposed drugs in two ways—we screen the most common comorbid proteins for their most frequent binding to repurposed drugs. For the set of comorbid diseases to a given complication, choose the drugs that treat the most complications."
It's critical to find the right drugs for those complications and side effects—and using the new "decision tool" can help do that, Skolnick says.
He cautions that researchers and clinicians must approach this objectively because "there are deep reasons why you have these 'off-target' interactions, where a drug binds to another protein that is not its original intended target—and that's what these algorithms take in to counter them. It's not magic" to run this methodology, Skolnick explains. "It's fractional assistance in a decision-making process—which comes with probabilities [that] it might work. If there's an 80 percent probability, you probably ought to try it."
More information: Courtney Astore et al, Prediction of severe adverse events, modes of action and drug treatments for COVID-19's complications, Scientific Reports (2021). DOI: 10.1038/s41598-021-00368-6
MOATAI-VIR methodology algorithms: sites.gatech.edu/cssb/moatai-vir/
Journal information: Scientific Reports
Provided by Georgia Institute of Technology