Researchers discover which molecules in blood serum can predict severity of COVID-19 infections
An estimated 14−17% of hospitalized COVID-19 patients develop a severe form of the disease, requiring oxygen support and admission to the intensive care unit. Underlying medical conditions such as diabetes, chronic cardiac diseases, chronic kidney diseases, obesity, and some genetic predispositions contribute to the severity of COVID-19. However, the severity of COVID-19 in a large group of patients cannot be explained only by these factors. At present, it is therefore very difficult to predict if the patient will develop a severe form of the disease. This is a problem, since such prediction and prognosis of the disease development from the first days is very important for providing adequate and timely management to reduce patients' mortality.
In a recent prospective clinical study, published in Frontiers of Immunology, an interdisciplinary team of immunologists, mathematicians, and clinicians from Belgium and Russia has identified a set of prediction markers which can be used to define the COVID-19 severity at the day of patient admission to the hospital. First author Dr. Olga Krysko (Ghent University): "The advantage of these identified markers is that these markers can be easily measured in most conventional clinical laboratories worldwide, making this prediction analysis accessible for many hospitals wordwide."
Artificial intelligence analysis
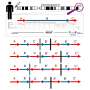
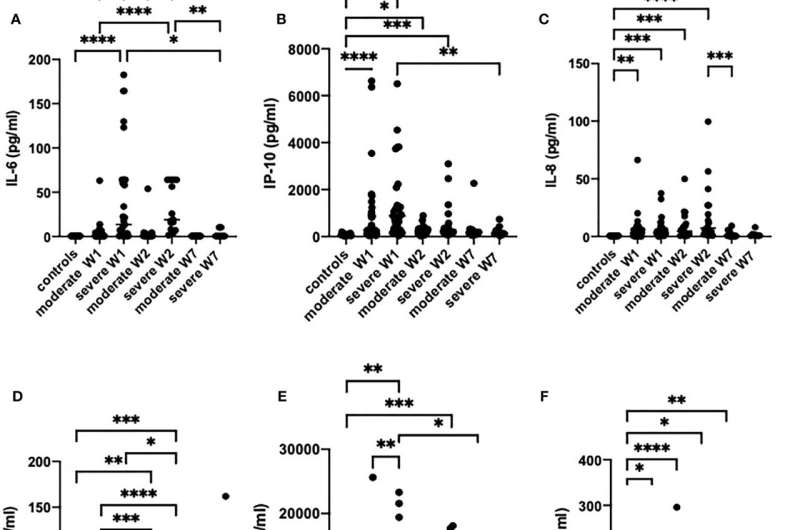
The prospective study analyzed 30 moderate and 30 severe COVID-19 patients and 17 healthy controls. Based on artificial intelligence analysis, eight parameters in blood serum (creatinine, glucose, monocyte number, fibrinogen, macrophage-derived cytokine, monokine induced by gamma interferon, C-reactive protein, and IL-6) have been shown to have a strong predictive value. Prof. Dr. Dmitri Krysko (a co-senior author of the study from Ghent University): "This methodology allows to make a prediction with an accuracy of about 83 to 87% whether the patient will develop a severe disease. These results pave the way for the development of new prognostic criteria and markers for COVID-19 patients and will help in the management of COVID-19 patients. We have developed an online tool, which is available for clinicians to calculate and estimate the prognosis for a particular patient. By entering the values of eight parameters the risk of COVID-19 severe is automatically calculated."
More information: Olga Krysko et al, Artificial Intelligence Predicts Severity of COVID-19 Based on Correlation of Exaggerated Monocyte Activation, Excessive Organ Damage and Hyperinflammatory Syndrome: A Prospective Clinical Study, Frontiers in Immunology (2021). DOI: 10.3389/fimmu.2021.715072