For the first time, scientists have shown that MRSA (methicillin-resistant Staphylococcus aureus) and other antibiotic-resistant 'superbug' infections can be tracked across Europe by combining whole-genome sequencing with a web-based system. In mBio today (May 5, 2016) researchers at Imperial College London and the Wellcome Trust Sanger Institute worked with a European network representing doctors in 450 hospitals in 25 countries to successfully interpret and visualise the spread of drug-resistant MRSA.
MRSA and other superbugs are a life-threatening problem for all hospitals across Europe with an estimated 400,000 cases per year and 25,000 deaths from resistant, hospital-acquired infections.
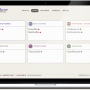
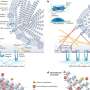
To enable infection control teams across Europe to easily share information and to form a dynamic picture of the rise and spread of antibiotic-resistant bacteria, the scientists from the newly formed Centre for Genomic Pathogen Surveillance developed Microreact.org, a web-based visualisation and mapping tool.
Dr David Aanensen, head of the Centre for Genomic Pathogen Surveillance and joint lead author on the paper said: "Drug resistance is a growing problem both in Europe and across the world and doctors need fast and accurate information to stop epidemics. Our study demonstrates the potential for combining whole-genome sequencing with internet-based visualisation tools to enable public health workers and doctors to see how an epidemic is spreading and make swift decisions to end it."
The research team read the whole genomes of S. aureus samples to identify which bugs are related to each other, and which are resistant to antibiotics. Using this approach, the scientists were not only able to show the rise and spread of MRSA across Europe, but also provide a quicker way to identify new hotspots of resistance.
Professor Hajo Grundmann, principal investigator on the study and Head of the Institute of Infection Prevention and Hospital Hygiene at the University Medical Centre Freiburg in Germany said: "One of the problems is that these bacteria not only spread within and between hospitals, but they also change their genetic properties due to evolutionary processes over time. Microreact.org allows us to look at their evolution within the context of how they are spreading across Europe."
In the paper, the scientists show that combining the drug-resistance profile of a bacteria with its whole-genome DNA sequence allowed them to build up a series of drug-resistance 'DNA photofits' for resistance to specific drugs. In the future, such an approach may help doctors decide on the best treatments more quickly and help bring drug-resistant outbreaks to an end.
Professor Ed Feil, joint lead author from the Milner Centre for Evolution, at the University of Bath, said: "We've developed user-friendly analysis software that demonstrates how whole genome sequence data can be a powerful tool for pan-European surveillance of MRSA and other important pathogens.
Being able to track the spread of outbreaks across the whole continent allows policymakers to identify potential risks to public health and implement appropriate prevention and control strategies."
More information: Aanensen D et al. Whole genome sequencing for routine pathogen surveillance in public health: A population snapshot of invasive Staphylococcus aureus in Europe. mBio 2016 Published on May 5, 2016. DOI: 10.1128/mBio.00444-16
Journal information: mBio
Provided by Wellcome Trust Sanger Institute