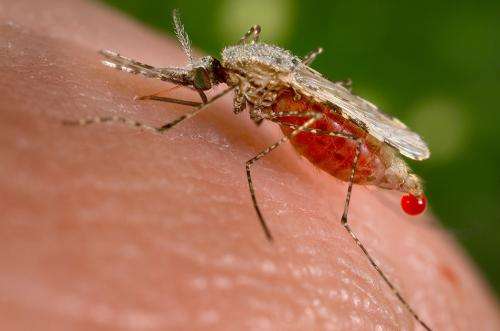
Credit: CDC
Scientists at the Wellcome Trust Sanger Institute and their collaborators have discovered genetic markers in malaria parasites linked with resistance to the anti-malarial drug piperaquine. Reported in Lancet Infectious Diseases, this research will allow health officials to monitor the spread of resistance, and help doctors and public health officers decide where the treatment is most likely to be effective.
Resistance to this key anti-malarial drug has recently emerged in Cambodia, leading to complete treatment failure there, threatening global efforts to treat and eliminate malaria.
Malaria is caused by Plasmodium parasites and in 2015, the World Health Organisation estimated that more than 200 million people were infected and nearly half a million people died worldwide from the disease. Children under the age of five made up 70 percent of these deaths. Malaria is a treatable disease when caught early enough, but is a huge problem in many areas due to drug resistance.
Piperaquine is a powerful drug, which is used in combination with another anti-malarial, artemisinin, as a first-line treatment in many areas of the world. Resistance to artemisinin emerged more than seven years ago in South East Asia, but until recently the combination of the drugs still successfully killed the malaria parasites there. Now, the development of piperaquine resistance has led to complete failure of treatment in Cambodia.
Researchers carried out a genome-wide association study on approximately 300 Plasmodium falciparum samples from Cambodia to study the genetic basis behind piperaquine resistance. They looked at thousands of variations in the DNA sequence of the parasites, comparing these across samples with different levels of resistance to piperaquine.
Dr Roberto Amato, lead author from the Sanger Institute, said: "By studying the genomes of these parasites we found two genetic markers that are linked with piperaquine resistance. Not only can we now use these markers to monitor the spread of the drug resistant malaria, they will also help towards understanding as much as possible about the biology and evolution of the parasite."
The scientists found that extra copies of the genes encoding two proteins of a family called plasmepsin, were linked with piperaquine resistance. Plasmepsins are part of a biological pathway that is targeted by other anti-malarial drugs, so this marker could also help the researchers understand the mechanism of the drug resistance. In addition to this, a mutation on chromosome 13 was found to be a second genetic marker linked with the resistance. Both markers were observed in parasites infecting patients who were not responding to treatment.
Dr Amato added: "The emergence of piperaquine resistance in these Cambodian parasites has led to complete treatment failure there. These malaria parasites are now resistant to both drugs, and since they are no longer being killed, resistance to both drugs will spread. This will threaten global attempts to eliminate malaria."
Professor Dominic Kwiatkowski, from the Sanger Institute said: "These findings provide the tools needed to map how far this resistance has spread, looking for these molecular markers in parasites in Cambodia and neighbouring countries. This will allow national malaria control programmes to rapidly recommend alternative therapies where possible and where needed, enhancing treatment for patients, and helping towards the ultimate goal of eliminating malaria."
More information: R Amato et al. Genetic markers associated with dihydroartemisinin-piperaquine failure in Plasmodium falciparum malaria in Cambodia: a genotype-phenotype association study, Lancet Infectious Diseases DOI: 10.1016/S1473-3099(16)30409-1 (2016).
Journal information: Lancet Infectious Diseases
Provided by Wellcome Trust Sanger Institute