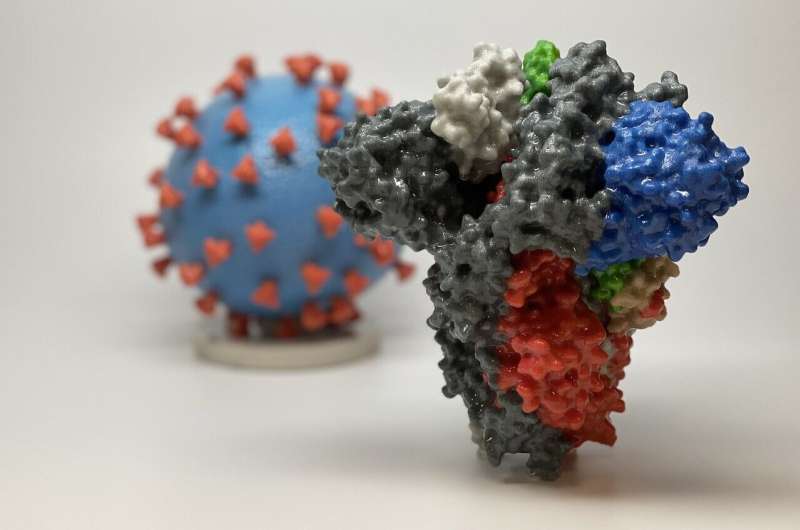
3D print of a spike protein of SARS-CoV-2, the virus that causes COVID-19—in front of a 3D print of a SARS-CoV-2 virus particle. The spike protein (foreground) enables the virus to enter and infect human cells. On the virus model, the virus surface (blue) is covered with spike proteins (red) that enable the virus to enter and infect human cells. Credit: NIH
Even as the first vaccines for SARS-CoV-2, the virus that causes COVID-19, are being distributed, scientists and clinicians around the world have remained steadfast in their efforts to better understand how the human immune system responds to the virus and protects people against it. Now, a research team—led by Johns Hopkins Medicine and in collaboration with ImmunoScape, a U.S.-Singapore biotechnology company—has published one of the most comprehensive characterizations to date of a critical contributor to that protection: the response of immune system cells called T lymphocytes (more commonly known as T cells) in people who have recovered from SARS-CoV-2 infection.
The researchers, whose findings were posted online Jan. 11, 2021, in The Journal of Clinical Investigation, say that better defining which T cells interact with which specific portions of the SARS-CoV-2 virus—as well as how those interactions can provide long-lasting immunity against COVID-19—may help spur development of the next generation of vaccines.
"We already knew that plasma from convalescent patients with COVID-19—those who have recovered from a SARS-CoV-2 infection—can contain antibodies that neutralize the virus, which can then be used to help other patients with active infection," says study senior author Thomas Quinn, M.D., professor of medicine at the Johns Hopkins University School of Medicine and National Institutes of Health distinguished investigator at the National Institute of Allergy and Infectious Diseases. "Our study was designed to assess which T cells react to specific proteins of SARS-CoV-2, how they might complement neutralizing antibodies in recovery from infection and what can be done to optimize the process for long-term protection."
The T cells of interest to Quinn and his colleagues are known as CD8+ T cells, also called cytotoxic or killer T cells for their ability to eliminate foreign invaders such as bacteria and viruses from the body. To analyze them, the researchers collected blood samples from 30 convalescent patients who had recovered from mild cases of COVID-19. The six human leukocyte antigens (HLAs, as they are more commonly known, are cell-surface proteins that regulate the immune system and are part of each person's genetic profile) of the donors studied, Quinn says, are representative of some 73% of the continental U.S. population, meaning the study results have broad significance.
The samples were taken from 26 to 62 days after the donors stopped having COVID-19 symptoms, so that "their immune response would be fully matured in response to the virus and have primed certain CD8+ T cells against it," says Quinn. The Johns Hopkins Medicine researchers measured the level of neutralizing antibodies in the donors at various times post-recovery and stored samples for deeper analysis.
That assessment occurred when the donor samples were sent to ImmunoScape for the difficult task of identifying which T cells had responded to SARS-CoV-2. More specifically, the company's deep immune cell profiling method could show toward which virus proteins the T cells directed that response—data that could provide valuable insight into the T cells' functional properties.
In a first-of-its-kind analysis, the ImmunoScape team used its highly sensitive HLA-SARS-CoV-2 tetramers—laboratory-produced proteins that bind exclusively to their T cell targets—to tag and identify the types of virus-recognizing CD8+ T cells. The samples were probed with 408 SARS-CoV-2 epitopes—proteins that may elicit an immune response—from the spikes on the virus surface, from the virus capsule and from nonstructural proteins inside the virus. The researchers then looked to see which T cells matched up with which epitopes.
"We found that 52 of the 408 epitopes were recognized by the T cells from the convalescent donors, with 18 of these epitope matchings previously unreported," says study co-author Aaron Tobian, M.D., Ph.D., director of the transfusion medicine division and professor of pathology at the Johns Hopkins University School of Medicine."Of these, 23% were derived from spike proteins—the types targeted by the currently available COVID-19 vaccines—while 14% were from capsule proteins and 63% were from nonstructural proteins that normally would not elicit an immune response."
Looking at the specific types of CD8+ T cells present at different times following recovery from COVID-19, the researchers found that as the levels of neutralizing antibodies increased in the convalescent plasma, so did the number of memory CD8+ T cells that recognized SARS-CoV-2 epitopes.
"That's good because those are the T cells you want to be primed in case you are exposed to SARS-CoV-2 a second time as they 'remember' the first infection and quickly direct the immune system to fight the virus before it takes root again," says Quinn.
Characterizing the CD8+ T cells in blood samples from convalescent patients that are specific to SARS-CoV-2 and, more importantly, specific to which viral epitopes, is an important achievement, Tobian adds.
"With this knowledge, we will be better equipped to design COVID-19 vaccines that produce a strong immune response and likely provide years of defense against SARS-CoV-2," he says.
More information: Hassen Kared et al. SARS-CoV-2-specific CD8+ T cell responses in convalescent COVID-19 individuals, Journal of Clinical Investigation (2021). DOI: 10.1172/JCI145476
Journal information: Journal of Clinical Investigation
Provided by Johns Hopkins University School of Medicine