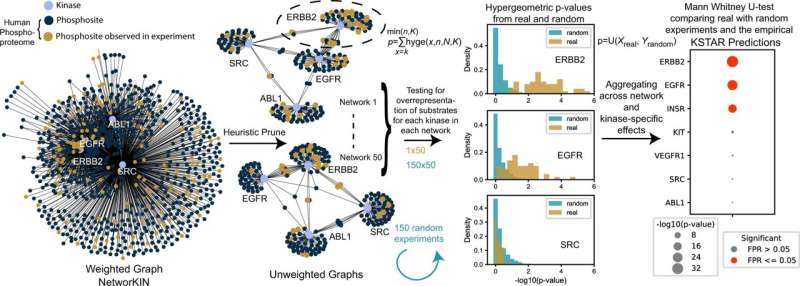
Overview of KSTAR algorithm. First, we heuristically prune dense and highly overlapping weighted kinase-substrate prediction graphs from NetworKIN into many sparse, binary graphs. Statistical enrichment is calculated for an experiment that has a defined set of phosphorylation sites for every kinase across all networks using a hypergeometric distribution. We generate and calculate enrichment in 150 random experiments using the same approach. Next, we use the Mann-Whitney U test to measure the likelihood that the enrichment p-values in the real experiment are more significant than the random experiments, giving us a final p-value, which accounts for the underlying enrichment of substrates in a network, aggregates that information across the different network configurations, and controls for the kinase- and experiment-specific behavior of enrichment that occurs by random chance. We measure the false positive rate by measuring the distribution-based test for a random experiment against the remaining 149 random experiments, repeating this for 100 times. Finally, the numerical KSTAR “score” (the -log10 transformation of the Mann–Whitney U-test) is presented in graphical format where the dot size is larger when there is more evidence phosphorylation sites are coordinately sampled from a kinase network. The FPR is indicated by “Significance” of having less than a specific empirical FPR. Credit: Nature Communications (2022). DOI: 10.1038/s41467-022-32017-5
UVA Health Cancer Center researchers have developed an algorithm that will improve cancer care by quickly and easily identifying patients who will benefit from powerful cancer drugs called kinase inhibitors. The algorithm may have other diagnostic benefits for patients as well.
Kinase inhibitors are the most common cancer drugs approved by the federal Food and Drug Administration. They can be hugely effective for the right patients, but they don't work for everyone. UVA's algorithm offers a new and better way to pinpoint patients who will benefit—an important step forward in precision medicine tailored to the individual.
"We are really excited about this algorithm, which performs better than existing approaches with fewer requirements and assumptions—making it more applicable to understanding a cancer state from a single snapshot of the tumor," said researcher Kristen M. Naegle of UVA's Department of Biomedical Engineering, a joint program of UVA's schools of Medicine and Engineering. "Combining this approach with existing biomarkers for cancer diagnosis may help us to better tailor therapies, design new combination therapies, anticipate response to treatment and design better clinical trials."
KSTAR for better cancer care
Naegle and colleagues set out to overcome the limitations of existing methods to identify patients who may benefit from kinase inhibitors. Most of these methods require difficult-to-obtain and sometimes unreliable information quantifying "phosphorylation sites" within cells. UVA's new approach, however, does not need all this complex measurement. Instead, it can predict key information based upon other available data.
This allows the algorithm to produce a specific "KSTAR score" for individual enzymes called kinases. Doctors can use these scores to determine which patients will respond to kinase inhibitors, helping guide the best treatment choices. Kinase inhibitors are widely used for certain types of blood, breast and lung cancers, among others. These drugs are often used in conjunction with other treatments, such as chemotherapy, radiation or surgery.
In testing their new algorithm, Naegle and her collaborators found that it worked reliably across different tissue types, suggesting it is useful for many types of cancer.
As an added benefit, KSTAR can serve as a diagnostic tool, the UVA researchers report. During their tests, KSTAR was able to determine that a patient with breast cancer was not HER2 positive, as doctors had previously believed. HER2-positive breast cancers can benefit from HER2-targeted kinase inhibition, but HER2-negative tumors will not, and HER2-inhibition comes with additional complications.
Additionally, the researchers find that around 20% of patients identified as HER2-negative by current clinical approaches actually have HER2 signatures that suggest they could benefit from HER2-targeted treatment—a treatment not currently offered to them. That type of information can be invaluable as doctors and patients discuss treatment options.
"We are collaborating and working with teams of researchers across a range of cancers to establish when and how KSTAR can help identify patient response to treatment," Naegle said. "We hope that we will be able to help better identify the right treatment for the right patient at the right time for better outcomes for patients."
Naegle and her collaborators have made their new algorithm freely available. They have published their findings in Nature Communications.
More information: Sam Crowl et al, KSTAR: An algorithm to predict patient-specific kinase activities from phosphoproteomic data, Nature Communications (2022). DOI: 10.1038/s41467-022-32017-5
Journal information: Nature Communications
Provided by University of Virginia