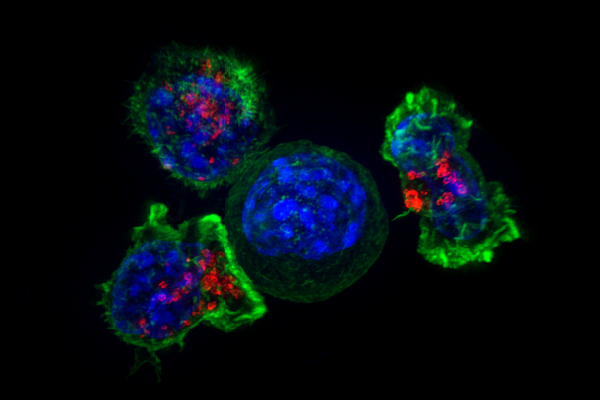
Killer T cells surround a cancer cell. Credit: NIH
(Medical Xpress)—A team of researchers from the Netherlands and the U.K. has developed a technique for studying inherited types of cancers using two relatively new technologies—organoid development and CRISPR/Cas9. In their paper published in the journal Science, the team describes their technique and how it can be used to better understand certain types of inherited cancers.
To better understand the factors at play during the formation of tumors due to inherited traits, the researchers created organoids from human intestinal stem cells. Organoids are organ-like constructs made of cells and tissue—they are grown artificially for experiments on living tissue that could not be done on actual organs. The organoids in this scenario were grown for genetic studies of tumors.
The study consisted of using CRISPR/Cas9 to cut a gene called NTHL1 from cells in the organoids. NTHL1 occurs naturally in the human colon. The point of cutting it from the DNA in the organoids was to find out if doing so resulted in a genetic signature similar to that of real cancer patients.
Under normal circumstances, NTHL1 creates proteins that are part of a process of excising defective genes. By removing the gene in the organoids, the researchers found that a mutational signature emerged that was similar to one found in a patient with a previously unknown type of breast cancer. A study of the genetic history of the patient revealed an inherited germline that led to inactivation of the NTHL1 gene, which in turn led to tumor growth. This meant that the technique had led to the identification of a gene defect related to a specific type of inheritable cancer.
By using the combined technologies to identify an underlying inheritable form of cancer, the researchers demonstrated that it is a viable research approach that could prove useful for better understanding the role that inherited DNA repair defects play in the development of inherited cancers. They suggest it could also be used to better understand the process involved in environmental mutagens, as well. They note, too, that their study shows that it is now possible to detect inheritable forms of cancers by sequencing the DNA of tumors that develop in cancer patients.
More information: Jarno Drost et al. Use of CRISPR-modified human stem cell organoids to study the origin of mutational signatures in cancer, Science (2017). DOI: 10.1126/science.aao3130
Abstract
Mutational processes underlie cancer initiation and progression. Signatures of these processes in cancer genomes may explain cancer etiology, and hold diagnostic and prognostic value. Here, we develop a strategy that can be used to explore the origin of cancer-associated mutational signatures. We used CRISPR/Cas9 technology to delete key DNA repair genes in human colon organoids, followed by delayed sub-cloning and whole-genome sequencing. We found that mutation accumulation in organoids deficient in the mismatch repair gene MLH1 is driven by replication errors and accurately models the mutation profiles observed in mismatch repair-deficient colorectal cancers. Application of this strategy to the cancer predisposition gene NTHL1, which encodes a base excision repair protein, revealed a mutational footprint (signature 30) previously observed in a breast cancer cohort. We show that signature 30 can arise from germline NTHL1 mutations.
Journal information: Science
© 2017 Medical Xpress