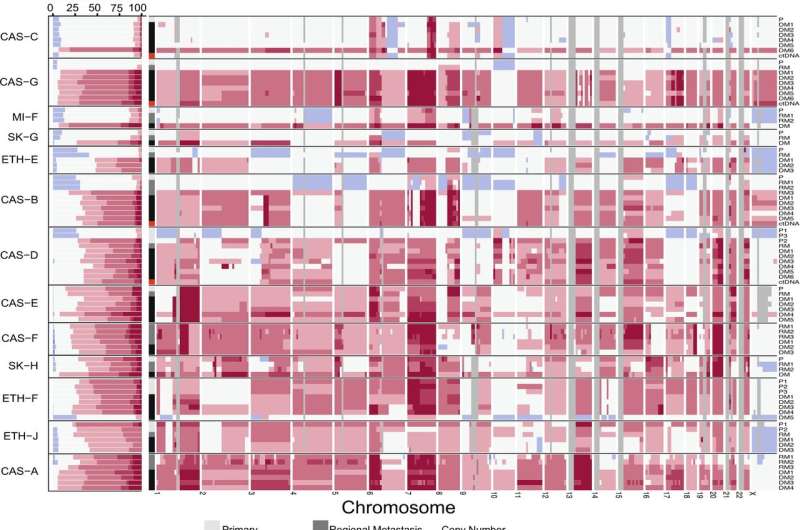
Ubiquitous hyperploidy in metastatic melanoma. A Left: genomic proportions of each tissue sample affected by copy number alterations (CNAs). Horizontal bars indicate the fraction of each genome with the indicated allelic copy number. Regions with total CN > 5 are represented by the same colour. Grey regions indicate undetermined CN. Right: CNAs in each sample at the cytogenetic band-level. B Validation of CN in 6 patients using Fluorescence In Situ Hybridisation (FISH). X-axis corresponds to the total CN of the segment overlapping the FISH probe predicted from sequencing data. Y-axis corresponds to the average CN across cells in each sample, determined by manual counting, for the indicated FISH probe. Centromeric probes corresponded to chromosomes 8, 9 and 11 as controls for probes for MYC, CDKN2A and CCND1, respectively, in those chromosomes. Cen: Centromeric probe. C FISH showing MYC amplification in two metastases from patient CAS-B, compared to the primary tumour (left). Arrow heads indicate cells with red (Myc) and green (centromere) signals. Scale bar = 10um. CTR: Centromere probes for chromosome 8. D Widespread genome doubling in melanoma. For each sample, the bar length indicates the Akaike Information Criteria (AIC) score obtained via branching process analyses of aneuploidy with genome doubling (left) versus aneuploidy alone (right). Solid circles indicate the most likely scenario for each sample.
Melbourne researchers have revealed how melanoma cells are flooded with DNA changes as this skin cancer progresses from early, treatable stages through to fatal end-stage disease.
Using genomics, the team tracked DNA changes occurring in melanoma samples donated by patients as their disease progressed, right through to the time the patient died. This revealed dramatic and chaotic genetic changes that accumulated in the melanoma cells as the cancers progressed, providing clues to potential new approaches to treating this disease.
The research, published in Nature Communications, was led by Professor Mark Shackleton, Professor Director of Oncology at Alfred Health and Monash University; Professor Tony Papenfuss, who leads WEHI's Computational Biology Theme and co-heads the Computational Cancer Biology Program at Peter MacCallum Cancer Centre; and Dr. Ismael Vergara, a computational biologist at WEHI, Peter Mac and the Melanoma Institute Australia.
Tracking a devastating cancer
Melanoma—the third most commonly diagnosed cancer in Australia—is caused by damaging changes occurring in the DNA of skin cells called melanocytes, usually as a result of exposure to ultraviolet (UV) radiation from sunlight. These genetic changes enable uncontrolled growth of the cells, forming a melanoma. As the melanoma cells keep dividing, some accumulate even more DNA changes, helping them to grow even faster and spread, said Professor Shackleton.
"At early stages, melanomas can be cured with surgery. However, they sometimes recur and progress to more aggressive forms. While there are excellent new therapies in these contexts, for some patients this progressing disease is difficult to treat," he said.
"We used DNA sequencing to document genetic changes that occurred as melanomas recurred and progressed in patients."
The team obtained genome sequencing data from tumors that had been donated by these patients and fed it into a mathematical model. This revealed that, as melanomas progress, they acquire increasingly dramatic genetic changes that add substantially to the initial DNA damage from UV radiation that caused the melanoma in the first place, said Professor Papenfuss.
"Early-stage primary melanomas showed changes in their DNA from UV damage—akin to mis-spelt words in a book. These changes were enough to allow the melanoma cells to grow uncontrollably in the skin," he said.
"In contrast, end-stage, highly aggressive melanomas, in addition to maintaining most of the original DNA damage, accumulated even more dramatic genetic changes. Every patient had melanoma cells in which the total amount of DNA had doubled—a very unusual phenomenon not seen in normal cells—but on top of that, large segments of DNA were rearranged or lost—like jumbled or missing pages in a book. We think this deluge of DNA changes 'supercharged' the genes that were driving the cancer, making the disease more aggressive.
"The genomes in the late-stage melanomas were completely chaotic. We think these mutations occur in a sudden, huge wave, distinct from to the gradual DNA changes that accumulate from UV exposure in form earlier-stage melanomas. The melanoma cells that acquire these chaotic changes seem to overwhelm the earlier, less-abnormal, slower growing cells," Professor Papenfuss said.
New insights into melanoma
Professor Shackleton said the research provided an in-depth explanation of how melanomas change as they grow and may also provide clues about how melanoma could be treated.
"We mapped sequential DNA changes to track the spread of the disease in individual cases, creating 'family trees' of melanoma cells that grew, spread and changed over time in each patient. In early-stage melanomas in the skin, the DNA changes were consistent with UV-damage, while the changes we saw in later-stage melanoma were totally wild, and associated with increased growth and spread of the disease, and evasion of the body's immune defenses. We could also link some DNA changes to the development of treatment resistance," he said.
The research also revealed key cancer genes that may contribute to the growth and spread of the melanoma.
"Many patients' late-stage melanomas had damage to genes known to control cell growth and to protect the structure of DNA during cell growth and division. When these genes don't work properly, cell growth becomes uncontrolled and the DNA inside cells becomes even more abnormal—it's a snowball effect. The findings also suggest that therapies which exploit these damaging changes might be useful for treating late-stage melanoma," Professor Shackleton said.
The study included tumor samples from Peter Mac's Cancer tissue Collection After Death (CASCADE) program—in which patients volunteer to undergo a rapid autopsy following their death.
"Our whole team would like to extend our sincere gratitude to the patients and their families whose participation in CASCADE made this research possible. We hope that the insights we have gained will lead to better treatments for people with melanoma," Professor Shackleton said.
More information: Ismael A. Vergara et al. Evolution of late-stage metastatic melanoma is dominated by aneuploidy and whole genome doubling, Nature Communications (2021). DOI: 10.1038/s41467-021-21576-8
Journal information: Nature Communications