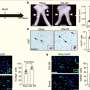
Scientists have discovered two separate genetic 'signatures' for prostate cancer that appear to be able to predict the severity of the disease, leading to hopes that in future, accuracy of prognosis and treatment of the disease could be greatly improved. Two Articles published in The Lancet Oncology reveal distinctive patterns of RNA—the genetic material that helps turn DNA into proteins—which appear to be able to predict whether patients have an aggressive prostate cancer, or whether they have a milder form of the disease.
Prostate cancer shows enormous variation between patients, with some people never showing symptoms, some responding well to treatment, and others developing resistance and progressing. Clinicians refer to the form of the disease which does not respond to standard androgen deprivation therapy as castration-resistant prostate cancer. Castration-resistant prostate cancer also shows wide variation in patient survival times, though the reasons for this are unclear.
Although tests to determine whether a patient has a more aggressive form of prostate cancer do currently exist, they are only moderately accurate. A more accurate way of determining whether a patient has a more dangerous form of prostate cancer would not only allow doctors to offer more accurate prognoses, but also enable better clinical trials for potential new treatments, as patients could be more effectively stratified into groups with aggressive or less aggressive disease.
The authors of one Article, led by Professor Johann de Bono at The Institute of Cancer Research, London, and The Royal Marsden NHS Foundation Trust in the UK, identified a set of genes which were able to predict whether patients had castration-resistant prostate cancer. Further, the signature stratified patients with castration-resistant prostate cancer: patients who were identified as having a distinctive nine-gene pattern characteristic of aggressive prostate cancer survived for an average of 9.2 months after referral for treatment, as opposed to 21.6 months in the group who did not test positive for the RNA signature under study.
In the other Article, researchers led by Professor William Oh at the Tisch Cancer Institute of Mount Sinai School of Medicine in the USA, identified a different set of genes with similar predictive properties to those identified by de Bono and colleagues. In this case, the researchers identified a set of six genes characteristic of a more aggressive form of prostate cancer, in a group of 62 patients at the Dana-Farber Cancer Institute in Boston. The signature divided patients into two groups: one with a median survival time of 7.8 months (the high-risk group), the other with a median survival of at least 34.9 months (the low-risk group). A validation cohort of 140 patients confirmed these findings.
Genetic signatures for different forms of cancer have been identified previously, but they have only been used for classification purposes – these studies are the first to show that they might have potential prognostic use. Previously, genetic tests like this were based on obtaining genetic material from the tumour itself, but these can be difficult to obtain and may not show the complete picture as there is evidence that patient prognosis may depend not only on the disease biology but also the individual's response to the disease. Furthermore, the genetic signatures identified from normal blood cells in these papers can be detected via a simple blood test, which could eventually lead to huge improvements in providing patients with prostate cancer with an accurate prognosis.
In a linked Comment, Dr Karina Dalsgaard Sørensen at Aarhus University Hospital in Denmark welcomes the findings, writing that, "Scarcity of prognostic markers presents a major challenge for the clinical management of castration-resistant prostate cancer. These results suggest that a few selected genes in blood samples from patients with castration-resistant prostate cancer can significantly improve the prediction of outcomes. However, the biological relevance of these prognostic signatures, which are the first of their kind, is largely unknown and further investigation into the underlying biological mechanisms at work here could greatly advance our understanding."
More information:
de Bono et al: www.thelancet.com/journals/lan … (12)70372-8/abstract
Oh et al: www.thelancet.com/journals/lan … (12)70263-2/abstract
Provided by Lancet